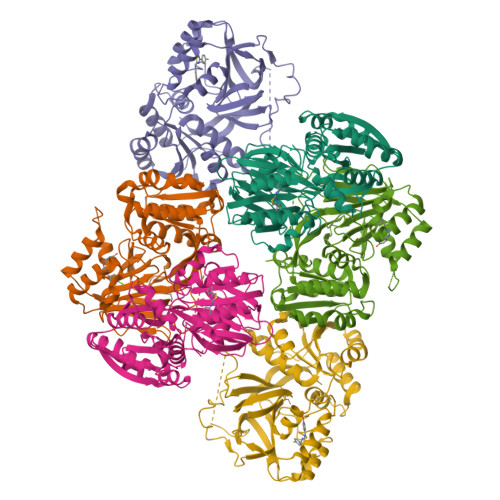
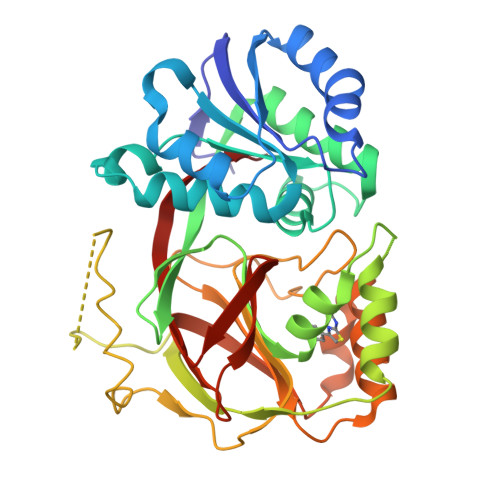
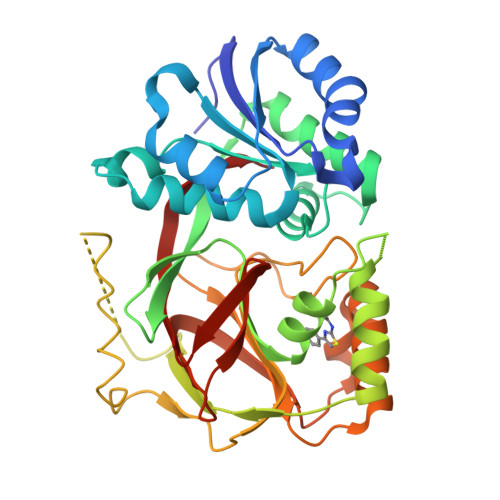
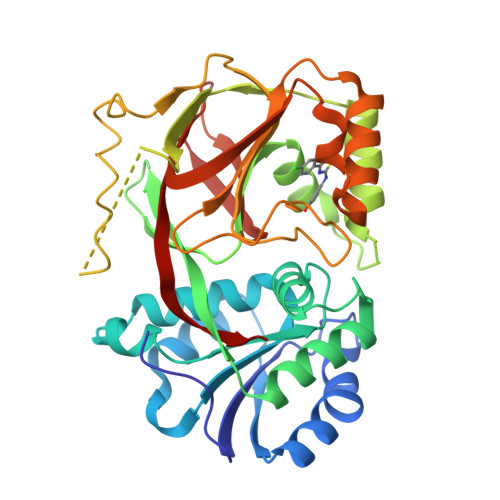
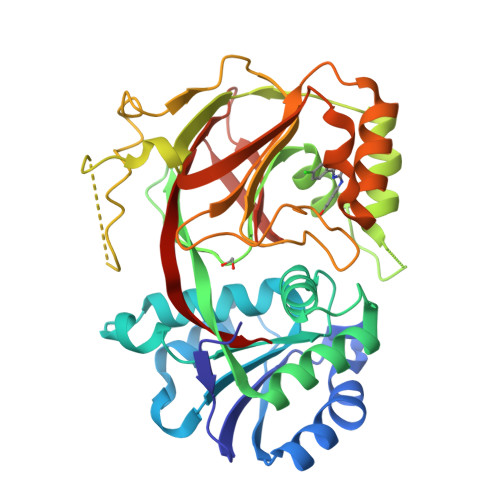
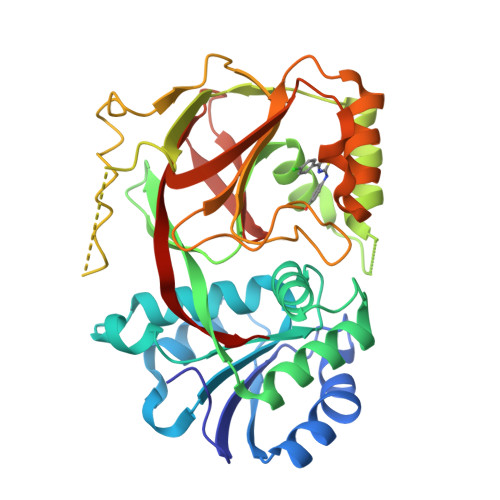
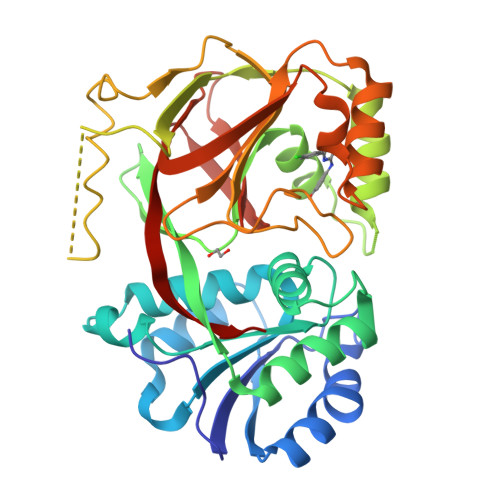
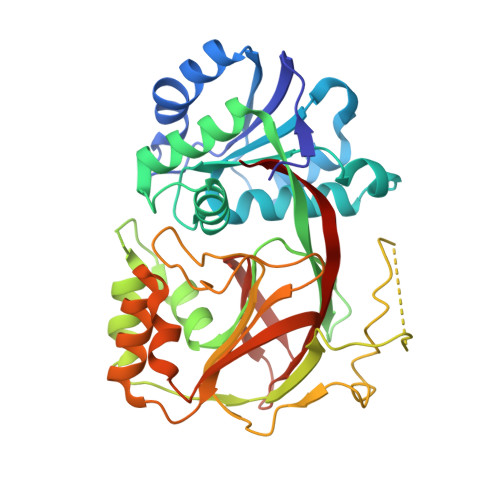
Molecular basis of sphingosine kinase 1 substrate recognition and catalysis.
Wang, Z., Min, X., Xiao, S.H., Johnstone, S., Romanow, W., Meininger, D., Xu, H., Liu, J., Dai, J., An, S., Thibault, S., Walker, N.(2013) Structure 21: 798-809
- PubMed: 23602659
- DOI: https://doi.org/10.1016/j.str.2013.02.025
- Primary Citation of Related Structures:
3VZB, 3VZC, 3VZD - PubMed Abstract:
Sphingosine kinase 1 (SphK1) is a lipid kinase that catalyzes the conversion of sphingosine to sphingosine-1-phosphate (S1P), which has been shown to play a role in lymphocyte trafficking, angiogenesis, and response to apoptotic stimuli. As a central enzyme in modulating the S1P levels in cells, SphK1 emerges as an important regulator for diverse cellular functions and a potential target for drug discovery. Here, we present the crystal structures of human SphK1 in the apo form and in complexes with a substrate sphingosine-like lipid, ADP, and an inhibitor at 2.0-2.3 Å resolution. The SphK1 structures reveal a two-domain architecture in which its catalytic site is located in the cleft between the two domains and a hydrophobic lipid-binding pocket is buried in the C-terminal domain. Comparative analysis of these structures with mutagenesis and kinetic studies provides insight into how SphK1 recognizes the lipid substrate and catalyzes ATP-dependent phosphorylation.
Organizational Affiliation:
Department of Molecular Structure and Characterization, Amgen, Inc., 1120 Veterans Boulevard, South San Francisco, CA 94080, USA. zwang@amgen.com