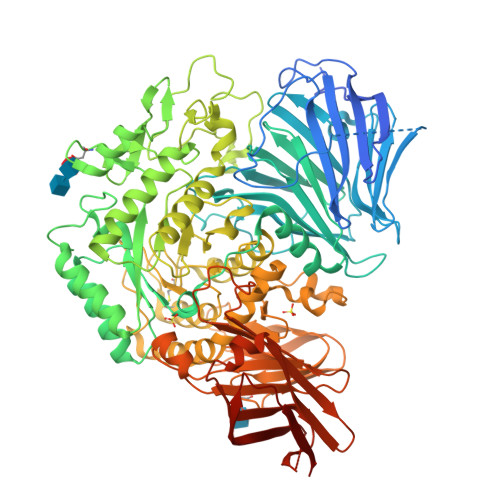
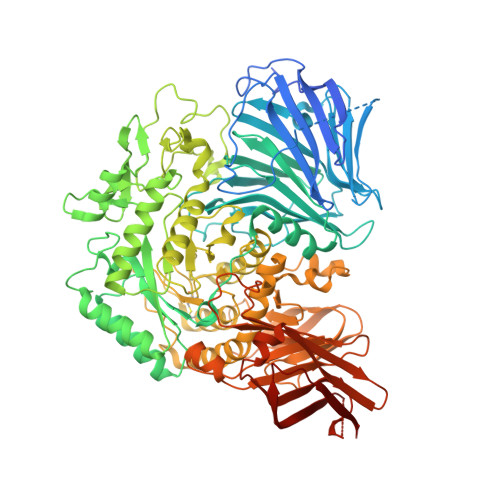
Molecular basis for the recognition of long-chain substrates by plant & alpha-glucosidase
Tagami, T., Yamashita, K., Okuyama, M., Mori, H., Yao, M., Kimura, A.(2013) J Biological Chem 288: 19296-19303
- PubMed: 23687304
- DOI: https://doi.org/10.1074/jbc.M113.465211
- Primary Citation of Related Structures:
3W37, 3W38 - PubMed Abstract:
Sugar beet α-glucosidase (SBG), a member of glycoside hydrolase family 31, shows exceptional long-chain specificity, exhibiting higher kcat/Km values for longer malto-oligosaccharides. However, its amino acid sequence is similar to those of other short chain-specific α-glucosidases. To gain structural insights into the long-chain substrate recognition of SBG, a crystal structure complex with the pseudotetrasaccharide acarbose was determined at 1.7 Å resolution. The active site pocket of SBG is formed by a (β/α)8 barrel domain and a long loop (N-loop) bulging from the N-terminal domain similar to other related enzymes. Two residues (Phe-236 and Asn-237) in the N-loop are important for the long-chain specificity. Kinetic analysis of an Asn-237 mutant enzyme and a previous study of a Phe-236 mutant enzyme demonstrated that these residues create subsites +2 and +3. The structure also indicates that Phe-236 and Asn-237 guide the reducing end of long substrates to subdomain b2, which is an additional element inserted into the (β/α)8 barrel domain. Subdomain b2 of SBG includes Ser-497, which was identified as the residue at subsite +4 by site-directed mutagenesis.
Organizational Affiliation:
Research Faculty of Agriculture, Hokkaido University, Sapporo 060-8589, Japan.