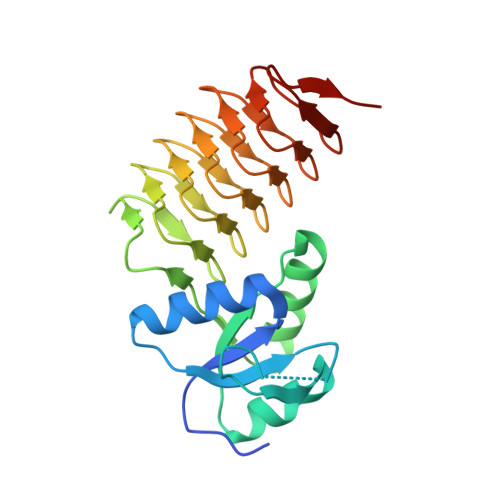
Catalytic Mechanism of Perosamine N-Acetyltransferase Revealed by High-Resolution X-ray Crystallographic Studies and Kinetic Analyses.
Thoden, J.B., Reinhardt, L.A., Cook, P.D., Menden, P., Cleland, W.W., Holden, H.M.(2012) Biochemistry 51: 3433-3444
- PubMed: 22443398
- DOI: https://doi.org/10.1021/bi300197h
- Primary Citation of Related Structures:
4EA7, 4EA8, 4EA9, 4EAA, 4EAB - PubMed Abstract:
N-Acetylperosamine is an unusual dideoxysugar found in the O-antigens of some Gram-negative bacteria, including the pathogenic Escherichia coli strain O157:H7. The last step in its biosynthesis is catalyzed by PerB, an N-acetyltransferase belonging to the left-handed β-helix superfamily of proteins. Here we describe a combined structural and functional investigation of PerB from Caulobacter crescentus. For this study, three structures were determined to 1.0 Å resolution or better: the enzyme in complex with CoA and GDP-perosamine, the protein with bound CoA and GDP-N-acetylperosamine, and the enzyme containing a tetrahedral transition state mimic bound in the active site. Each subunit of the trimeric enzyme folds into two distinct regions. The N-terminal domain is globular and dominated by a six-stranded mainly parallel β-sheet. It provides most of the interactions between the protein and GDP-perosamine. The C-terminal domain consists of a left-handed β-helix, which has nearly seven turns. This region provides the scaffold for CoA binding. On the basis of these high-resolution structures, site-directed mutant proteins were constructed to test the roles of His 141 and Asp 142 in the catalytic mechanism. Kinetic data and pH-rate profiles are indicative of His 141 serving as a general base. In addition, the backbone amide group of Gly 159 provides an oxyanion hole for stabilization of the tetrahedral transition state. The pH-rate profiles are also consistent with the GDP-linked amino sugar substrate entering the active site in its unprotonated form. Finally, for this investigation, we show that PerB can accept GDP-3-deoxyperosamine as an alternative substrate, thus representing the production of a novel trideoxysugar.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin, Madison, WI 53706, USA.