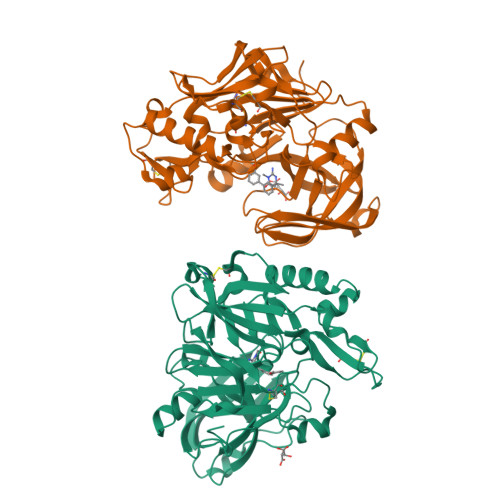
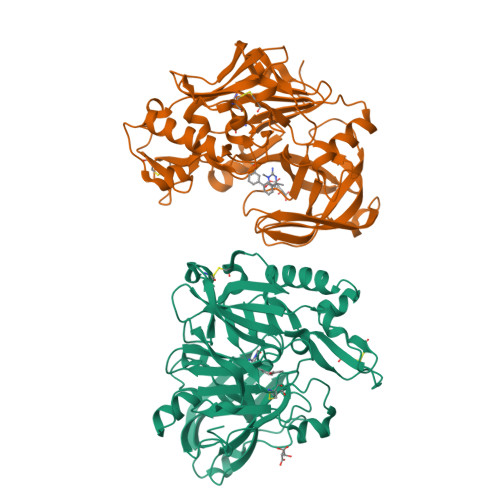
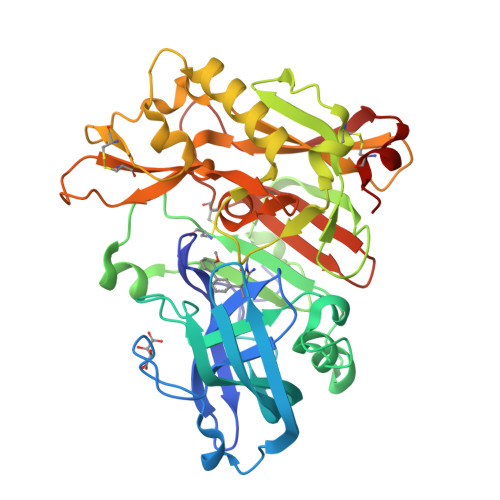
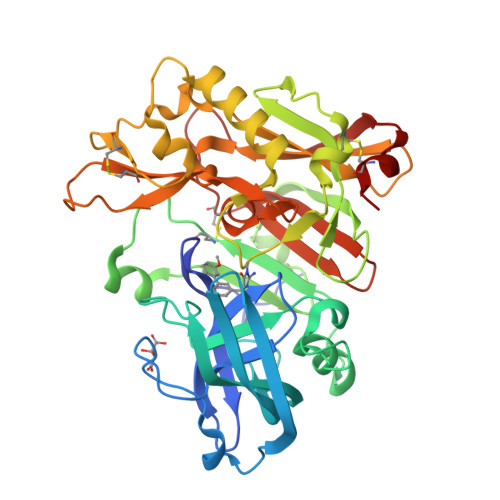
A Potent and Orally Efficacious, Hydroxyethylamine-Based Inhibitor of beta-Secretase.
Kaller, M.R., Harried, S.S., Albrecht, B., Amarante, P., Babu-Khan, S., Bartberger, M.D., Brown, J., Brown, R., Chen, K., Cheng, Y., Citron, M., Croghan, M.D., Graceffa, R., Hickman, D., Judd, T., Kriemen, C., La, D., Li, V., Lopez, P., Luo, Y., Masse, C., Monenschein, H., Nguyen, T., Pennington, L.D., Miguel, T.S., Sickmier, E.A., Wahl, R.C., Weiss, M.M., Wen, P.H., Williamson, T., Wood, S., Xue, M., Yang, B., Zhang, J., Patel, V., Zhong, W., Hitchcock, S.(2012) ACS Med Chem Lett 3: 886-891
- PubMed: 24900403
- DOI: https://doi.org/10.1021/ml3000148
- Primary Citation of Related Structures:
4DUS, 4FS4 - PubMed Abstract:
β-Secretase inhibitors are potentially disease-modifying treatments for Alzheimer's disease. Previous efforts in our laboratory have resulted in hydroxyethylamine-derived inhibitors such as 1 with low nanomolar potency against β-site amyloid precursor protein cleaving enzyme (BACE). When dosed intravenously, compound 1 was also shown to significantly reduce Aβ40 levels in plasma, brain, and cerebral spinal fluid. Herein, we report further optimizations that led to the discovery of inhibitor 16 as a novel, potent, and orally efficacious BACE inhibitor.
Organizational Affiliation:
Chemistry Research and Discovery, Department of Molecular Structure, Department of Neuroscience, Department of HTS and Molecular Pharmacology, and Department of Pharmacokinetics and Drug Metabolism, Amgen Inc. , One Amgen Center Drive, Thousand Oaks, California 91320, United States.