P2' benzene carboxylic acid moiety is associated with decrease in cellular uptake: evaluation of novel non-peptidic HIV-1 protease inhibitors containing P2 bis-tetrahydrofuran moiety.
Yedidi, R.S., Maeda, K., Fyvie, W.S., Steffey, M., Davis, D.A., Palmer, I., Aoki, M., Kaufman, J.D., Stahl, S.J., Garimella, H., Das, D., Wingfield, P.T., Ghosh, A.K., Mitsuya, H.(2013) Antimicrob Agents Chemother 57: 4920-4927
- PubMed: 23877703
- DOI: https://doi.org/10.1128/AAC.00868-13
- Primary Citation of Related Structures:
4HLA, 4I8W, 4I8Z - PubMed Abstract:
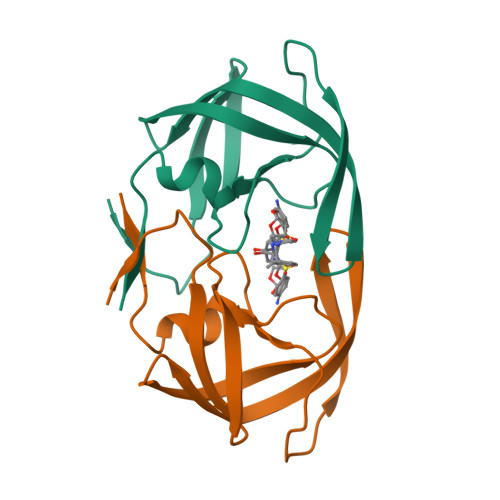
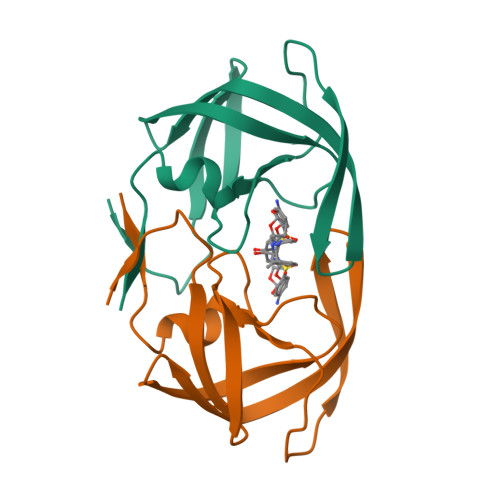
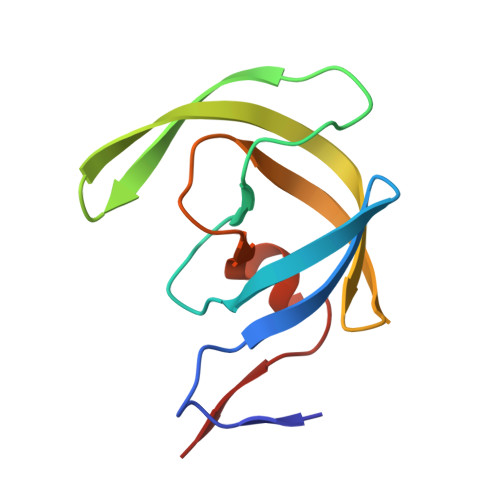
GRL007 and GRL008, two structurally related nonpeptidic human immunodeficiency virus type 1 (HIV-1) protease inhibitors (PIs) containing 3(R),3a(S),6a(R)-bis-tetrahydrofuranylurethane (bis-THF) as the P2 moiety and a sulfonamide isostere consisting of benzene carboxylic acid and benzene carboxamide as the P2' moiety, respectively, were evaluated for their antiviral activity and interactions with wild-type protease (PR(WT)). Both GRL007 (Ki of 12.7 pM with PR(WT)) and GRL008 (Ki of 8.9 pM) inhibited PR(WT) with high potency in vitro. X-ray crystallographic analysis of PR(WT) in complex with GRL007 or GRL008 showed that the bis-THF moiety of both compounds has three direct polar contacts with the backbone amide nitrogen atoms of Asp29 and Asp30 of PR(WT). The P2' moiety of both compounds showed one direct contact with the backbone of Asp30' and a bridging polar contact with Gly48' through a water molecule. Cell-based antiviral assays showed that GRL007 was inactive (50% effective concentration [EC50] of >1 μM) while GRL008 was highly active (EC50 of 0.04 μM) against wild-type HIV-1. High-performance liquid chromatography (HPLC)/mass spectrometry-based cellular uptake assays showed 8.1- and 84-fold higher intracellular concentrations of GRL008 than GRL007 in human MT-2 and MT-4 cell extracts, respectively. Thus, GRL007, in spite of its favorable enzyme-inhibitory activity and protease binding profile, exhibited a lack of antiviral activity in cell-based assays, most likely due to its compromised cellular uptake associated with its P2' benzene carboxylic acid moiety. The anti-HIV-1 potency, favorable toxicity, and binding profile of GRL008 suggest that further optimization of the P2' moiety may improve its antiretroviral features.
Organizational Affiliation:
Experimental Retrovirology Section.