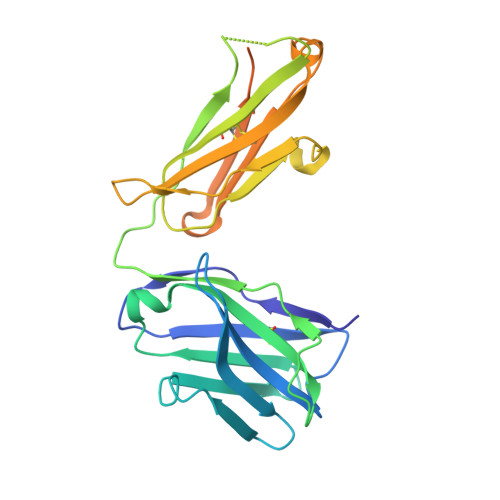
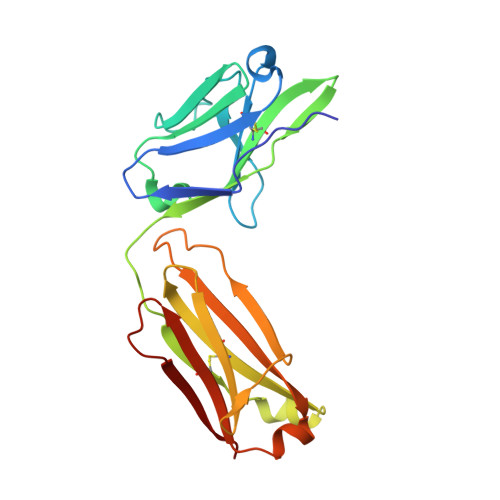
Structural basis of HCV neutralization by human monoclonal antibodies resistant to viral neutralization escape.
Krey, T., Meola, A., Keck, Z.Y., Damier-Piolle, L., Foung, S.K., Rey, F.A.(2013) PLoS Pathog 9: e1003364-e1003364
- PubMed: 23696737
- DOI: https://doi.org/10.1371/journal.ppat.1003364
- Primary Citation of Related Structures:
4JZN, 4JZO - PubMed Abstract:
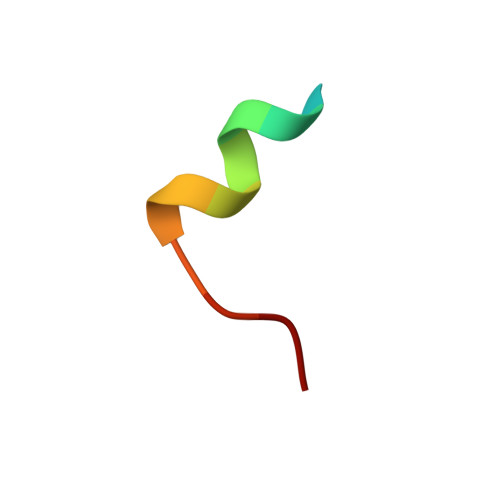
The high mutation rate of hepatitis C virus allows it to rapidly evade the humoral immune response. However, certain epitopes in the envelope glycoproteins cannot vary without compromising virus viability. Antibodies targeting these epitopes are resistant to viral escape from neutralization and understanding their binding-mode is important for vaccine design. Human monoclonal antibodies HC84-1 and HC84-27 target conformational epitopes overlapping the CD81 receptor-binding site, formed by segments aa434-446 and aa610-619 within the major HCV glycoprotein E2. No neutralization escape was yet observed for these antibodies. We report here the crystal structures of their Fab fragments in complex with a synthetic peptide comprising aa434-446. The structures show that the peptide adopts an α-helical conformation with the main contact residues F⁴⁴² and Y⁴⁴³ forming a hydrophobic protrusion. The peptide retained its conformation in both complexes, independently of crystal packing, indicating that it reflects a surface feature of the folded glycoprotein that is exposed similarly on the virion. The same residues of E2 are also involved in interaction with CD81, suggesting that the cellular receptor binds the same surface feature and potential escape mutants critically compromise receptor binding. In summary, our results identify a critical structural motif at the E2 surface, which is essential for virus propagation and therefore represents an ideal candidate for structure-based immunogen design for vaccine development.
Organizational Affiliation:
Institut Pasteur, Unité de Virologie Structurale, Departement Virologie, Paris, France. tkrey@pasteur.fr