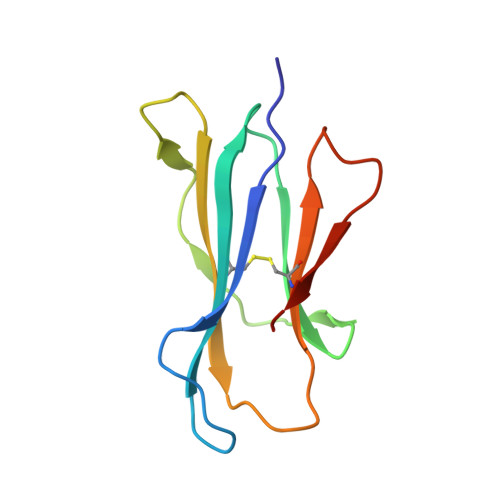
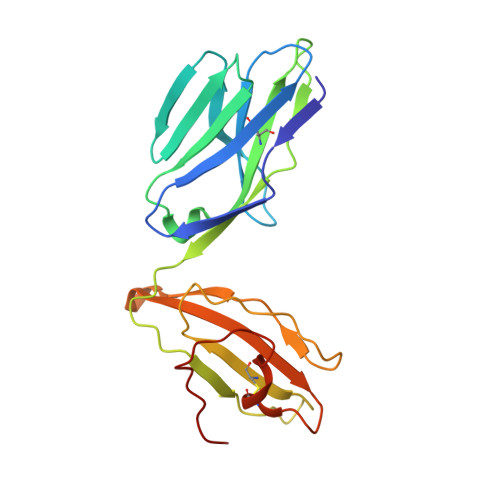
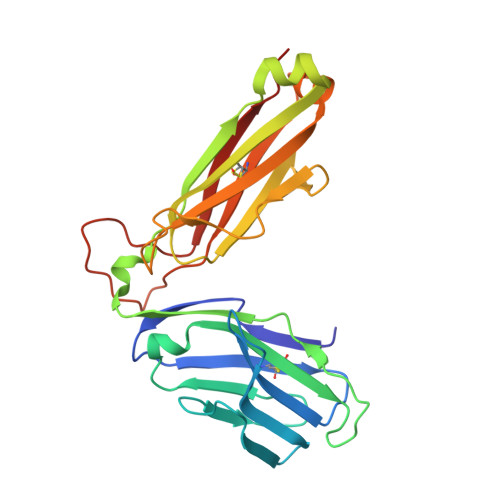
Recognition of vitamin B metabolites by mucosal-associated invariant T cells.
Patel, O., Kjer-Nielsen, L., Le Nours, J., Eckle, S.B., Birkinshaw, R., Beddoe, T., Corbett, A.J., Liu, L., Miles, J.J., Meehan, B., Reantragoon, R., Sandoval-Romero, M.L., Sullivan, L.C., Brooks, A.G., Chen, Z., Fairlie, D.P., McCluskey, J., Rossjohn, J.(2013) Nat Commun 4: 2142-2142
- PubMed: 23846752
- DOI: https://doi.org/10.1038/ncomms3142
- Primary Citation of Related Structures:
4L4T, 4L4V - PubMed Abstract:
The mucosal-associated invariant T-cell antigen receptor (MAIT TCR) recognizes MR1 presenting vitamin B metabolites. Here we describe the structures of a human MAIT TCR in complex with human MR1 presenting a non-stimulatory ligand derived from folic acid and an agonist ligand derived from a riboflavin metabolite. For both vitamin B antigens, the MAIT TCR docks in a conserved manner above MR1, thus acting as an innate-like pattern recognition receptor. The invariant MAIT TCR α-chain usage is attributable to MR1-mediated interactions that prise open the MR1 cleft to allow contact with the vitamin B metabolite. Although the non-stimulatory antigen does not contact the MAIT TCR, the stimulatory antigen does. This results in a higher affinity of the MAIT TCR for a stimulatory antigen in comparison with a non-stimulatory antigen. We formally demonstrate a structural basis for MAIT TCR recognition of vitamin B metabolites, while illuminating how TCRs recognize microbial metabolic signatures.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Clayton, Victoria 3800, Australia.