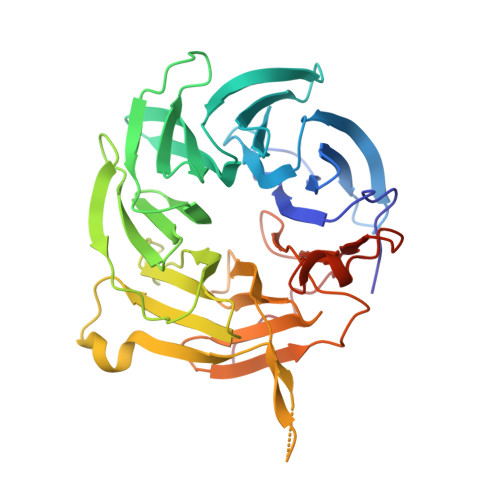
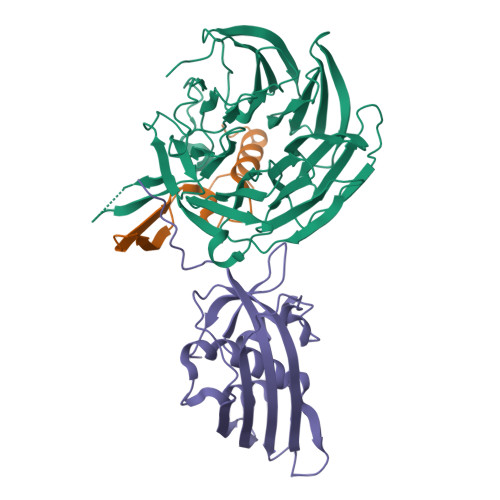
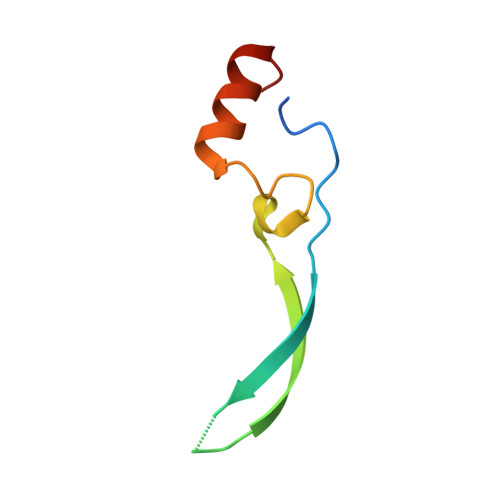
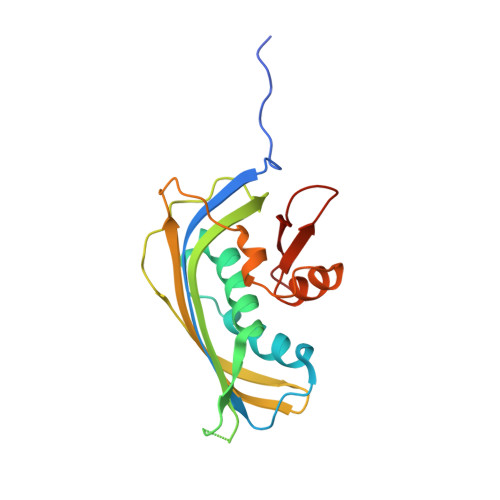
Vesiculoviral matrix (M) protein occupies nucleic acid binding site at nucleoporin pair (Rae1 Nup98).
Quan, B., Seo, H.S., Blobel, G., Ren, Y.(2014) Proc Natl Acad Sci U S A 111: 9127-9132
- PubMed: 24927547
- DOI: https://doi.org/10.1073/pnas.1409076111
- Primary Citation of Related Structures:
4OWR - PubMed Abstract:
mRNA export factor 1 (Rae1) and nucleoporin 98 (Nup98) are host cell targets for the matrix (M) protein of vesicular stomatitis virus (VSV). How Rae1 functions in mRNA export and how M protein targets both Rae1 and Nup98 are not understood at the molecular level. To obtain structural insights, we assembled a 1:1:1 complex of M•Rae1•Nup98 and established a crystal structure at 3.15-Å resolution. We found that the M protein contacts the Rae1•Nup98 heterodimer principally by two protrusions projecting from the globular domain of M like a finger and thumb. Both projections clamp to the side of the β-propeller of Rae1, with the finger also contacting Nup98. The most prominent feature of the finger is highly conserved Methionine 51 (Met51) with upstream and downstream acidic residues. The complementary surface on Rae1 displays a deep hydrophobic pocket, into which Met51 fastens like a bolt, and a groove of basic residues on either side, which bond to the acidic residues of the finger. Notably, the M protein competed for in vitro binding of various oligonucleotides to Rae1•Nup98. We localized this competing activity of M to its finger using a synthetic peptide. Collectively, our data suggest that Rae1 serves as a binding protein for the phosphate backbone of any nucleic acid and that the finger of M mimics this ligand. In the context of mRNA export, we propose that a given mRNA segment, after having been deproteinated by helicase, is transiently reproteinated by Nup98-tethered Rae1. We suggest that such repetitive cycles provide cytoplasmic stopover sites required for ratcheting mRNA across the nuclear pore.
Organizational Affiliation:
Laboratory of Cell Biology and Howard Hughes Medical Institute, The Rockefeller University, New York, NY 10065.