Unraveling the B.Pseudomallei Heptokinase Wcbl: From Structure to Drug Discovery.
Vivoli, M., Isupov, M.N., Nicholas, R., Hill, A., Scott, A.E., Kosma, P., Prior, J.L., Harmer, N.J.(2015) Chem Biol 22: 1622
- PubMed: 26687481
- DOI: https://doi.org/10.1016/j.chembiol.2015.10.015
- Primary Citation of Related Structures:
4USK, 4USM, 4UT4, 4UTG - PubMed Abstract:
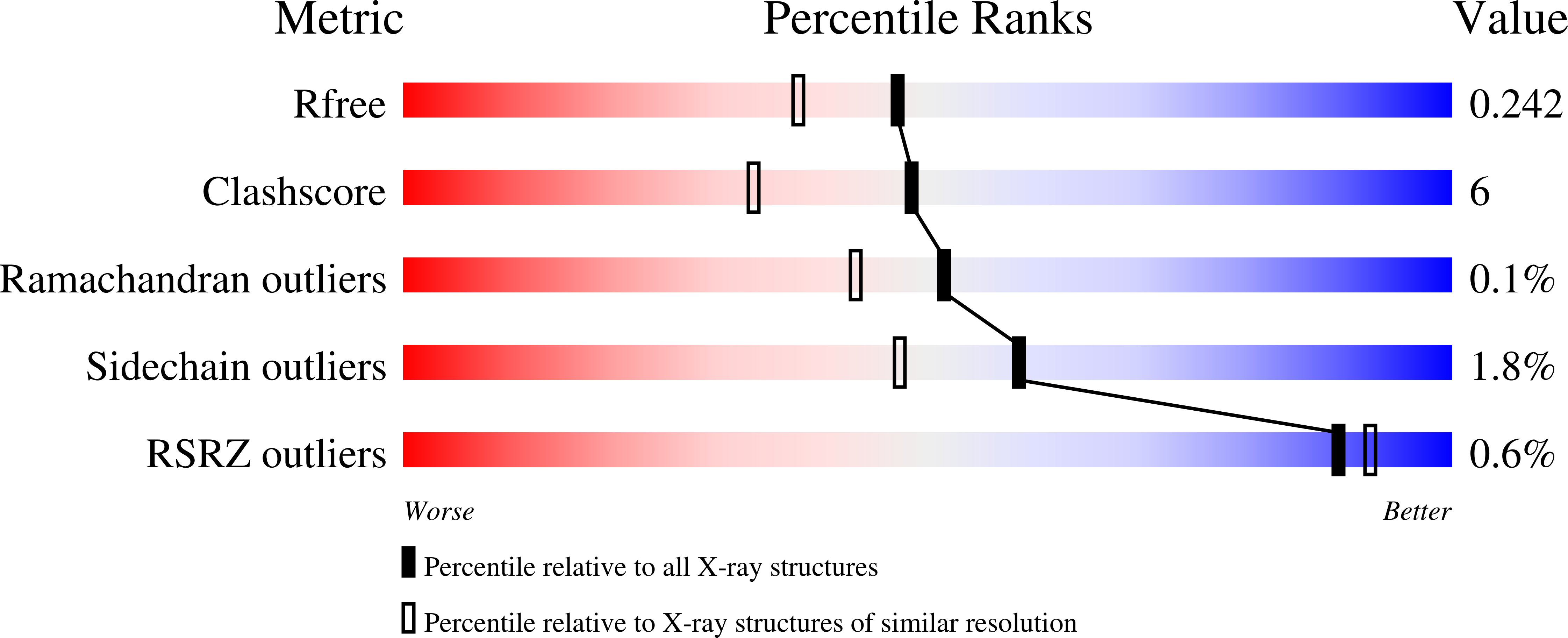
Gram-negative bacteria utilize heptoses as part of their repertoire of extracellular polysaccharide virulence determinants. Disruption of heptose biosynthesis offers an attractive target for novel antimicrobials. A critical step in the synthesis of heptoses is their 1-O phosphorylation, mediated by kinases such as HldE or WcbL. Here, we present the structure of WcbL from Burkholderia pseudomallei. We report that WcbL operates through a sequential ordered Bi-Bi mechanism, loading the heptose first and then ATP. We show that dimeric WcbL binds ATP anti-cooperatively in the absence of heptose, and cooperatively in its presence. Modeling of WcbL suggests that heptose binding causes an elegant switch in the hydrogen-bonding network, facilitating the binding of a second ATP molecule. Finally, we screened a library of drug-like fragments, identifying hits that potently inhibit WcbL. Our results provide a novel mechanism for control of substrate binding and emphasize WcbL as an attractive anti-microbial target for Gram-negative bacteria.
Organizational Affiliation:
Department of Biosciences, University of Exeter, Henry Wellcome Building, Stocker Road, Exeter EX4 4QD, UK.