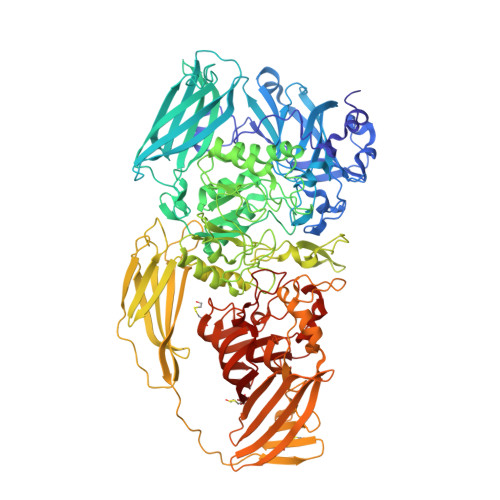
High resolution refinement of beta-galactosidase in a new crystal form reveals multiple metal-binding sites and provides a structural basis for alpha-complementation.
Juers, D.H., Jacobson, R.H., Wigley, D., Zhang, X.J., Huber, R.E., Tronrud, D.E., Matthews, B.W.(2000) Protein Sci 9: 1685-1699
- PubMed: 11045615
- DOI: https://doi.org/10.1110/ps.9.9.1685
- Primary Citation of Related Structures:
1DP0, 1F4A, 1F4H, 4V41 - PubMed Abstract:
The unrefined fold of Escherichia coli beta-galactosidase based on a monoclinic crystal form with four independent tetramers has been reported previously. Here, we describe a new, orthorhombic form with one tetramer per asymmetric unit that has permitted refinement of the structure at 1.7 A resolution. This high-resolution analysis has confirmed the original description of the structure and revealed new details. An essential magnesium ion, identified at the active site in the monoclinic crystals, is also seen in the orthorhombic form. Additional putative magnesium binding sites are also seen. Sodium ions are also known to affect catalysis, and five putative binding sites have been identified, one close to the active site. In a crevice on the protein surface, five linked five-membered solvent rings form a partial clathrate-like structure. Some other unusual aspects of the structure include seven apparent cis-peptide bonds, four of which are proline, and several internal salt-bridge networks. Deep solvent-filled channels and tunnels extend across the surface of the molecule and pass through the center of the tetramer. Because of these departures from a compact globular shape, the molecule is not well characterized by prior empirical relationships between the mass and surface area of proteins. The 50 or so residues at the amino terminus have a largely extended conformation and mostly lie across the surface of the protein. At the same time, however, segment 13-21 contributes to a subunit interface, and residues 29-33 pass through a "tunnel" formed by a domain interface. Taken together, the overall arrangement provides a structural basis for the phenomenon of alpha-complementation.
Organizational Affiliation:
Institute of Molecular Biology, and Department of Physics, University of Oregon, Eugene 97403-1229, USA.