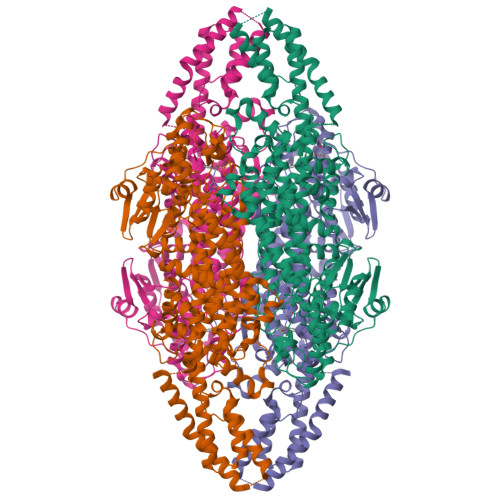
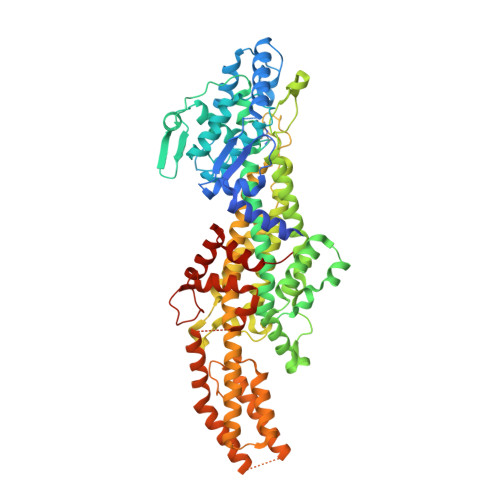
Structural Investigations Into the Stereochemistry and Activity of a Phenylalanine-2,3-Aminomutase from Taxus Chinensis.
Wybenga, G.G., Szymanski, W., Wu, B., Feringa, B.L., Janssen, D.B., Dijkstra, B.W.(2014) Biochemistry 53: 3187
- PubMed: 24786474
- DOI: https://doi.org/10.1021/bi500187a
- Primary Citation of Related Structures:
4C5R, 4C5S, 4C5U, 4C6G, 4CQ5 - PubMed Abstract:
Phenylalanine-2,3-aminomutase (PAM) from Taxus chinensis, a 4-methylidene-imidazole-5-one (MIO)-dependent enzyme, catalyzes the reversible conversion of (S)-α-phenylalanine into (R)-β-phenylalanine via trans-cinnamic acid. The enzyme also catalyzes the direct addition of ammonia to trans-cinnamic acid, a reaction that can be used for the preparation of β-amino acids, which occur as frequent constituents of bioactive compounds. Different hypotheses have been formulated to explain the stereochemistry of the PAM-catalyzed reaction, but structural evidence for these hypotheses is lacking. Furthermore, it remains unclear how the PAM MIO group is formed from the three-amino acid (A-S-G) sequence motif. For these reasons, we elucidated PAM three-dimensional (3D) structures with a bound (R)-β-phenylalanine analogue and with bound trans-cinnamic acid. In addition, 3D structures of the (inactive) Y322A and N231A mutants of PAM were elucidated, which were found to be MIO-less. We conclude that the stereochemistry of the PAM-catalyzed reaction originates from the enzyme's ability to bind trans-cinnamic acid in two different orientations, with either the si,si face or the re,re face directed toward the MIO group, as evidenced by two distinct carboxylate binding modes. The results also suggest that the N231 side chain promotes MIO group formation by increasing the nucleophilicity of the G177 N atom through acidification of the amide proton.
Organizational Affiliation:
Laboratory of Biophysical Chemistry, University of Groningen , Nijenborgh 7, 9747 AG Groningen, The Netherlands.