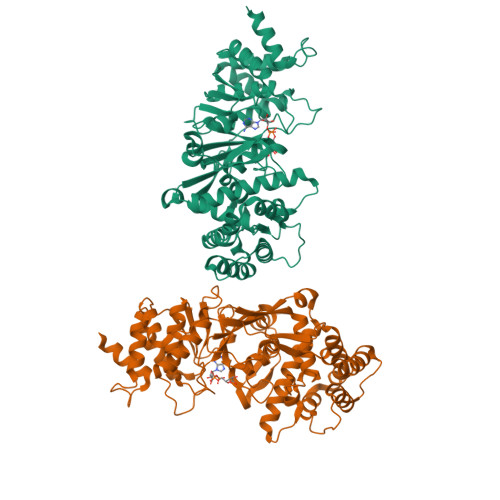
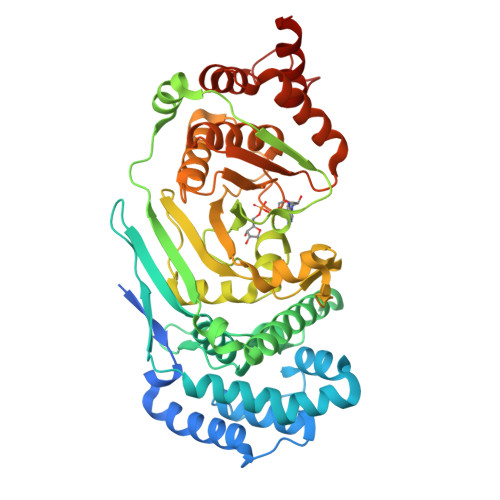
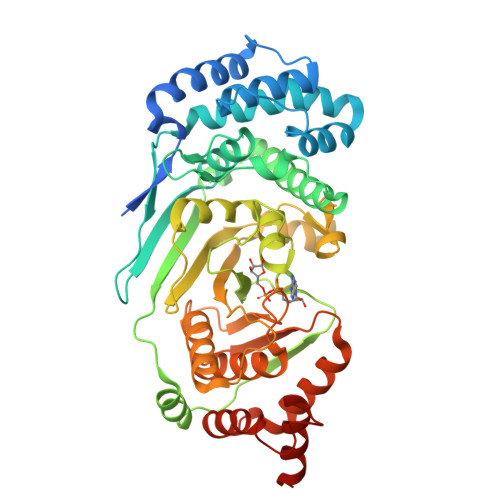
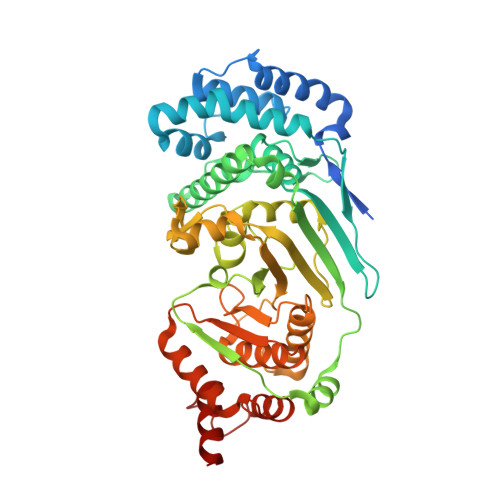
Structure and mechanism of a canonical poly(ADP-ribose) glycohydrolase.
Dunstan, M.S., Barkauskaite, E., Lafite, P., Knezevic, C.E., Brassington, A., Ahel, M., Hergenrother, P.J., Leys, D., Ahel, I.(2012) Nat Commun 3: 878-878
- PubMed: 22673905
- DOI: https://doi.org/10.1038/ncomms1889
- Primary Citation of Related Structures:
4EPP, 4EPQ - PubMed Abstract:
Poly(ADP-ribosyl)ation is a reversible post-translational protein modification involved in the regulation of a number of cellular processes including DNA repair, chromatin structure, mitosis, transcription, checkpoint activation, apoptosis and asexual development. The reversion of poly(ADP-ribosyl)ation is catalysed by poly(ADP-ribose) (PAR) glycohydrolase (PARG), which specifically targets the unique PAR (1''-2') ribose-ribose bonds. Here we report the structure and mechanism of the first canonical PARG from the protozoan Tetrahymena thermophila. In addition, we reveal the structure of T. thermophila PARG in a complex with a novel rhodanine-containing mammalian PARG inhibitor RBPI-3. Our data demonstrate that the protozoan PARG represents a good model for human PARG and is therefore likely to prove useful in guiding structure-based discovery of new classes of PARG inhibitors.
Organizational Affiliation:
Manchester Interdisciplinary Biocentre, Princess Street 131, M1 7DN, Manchester, UK.