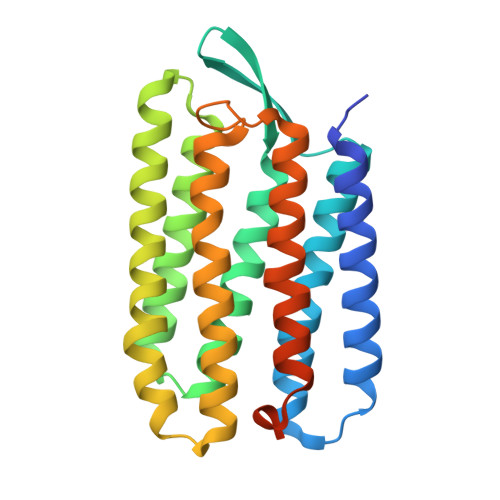
Membrane protein structure determination by SAD, SIR, or SIRAS phasing in serial femtosecond crystallography using an iododetergent
Nakane, T., Hanashima, S., Suzuki, M., Saiki, H., Hayashi, T., Kakinouchi, K., Sugiyama, S., Kawatake, S., Matsuoka, S., Matsumori, N., Nango, E., Kobayashi, J., Shimamura, T., Kimura, K., Mori, C., Kunishima, N., Sugahara, M., Takakyu, Y., Inoue, S., Masuda, T., Hosaka, T., Tono, K., Joti, Y., Kameshima, T., Hatsui, T., Yabashi, M., Inoue, T., Nureki, O., Iwata, S., Murata, M., Mizohata, E.(2016) Proc Natl Acad Sci U S A 113: 13039-13044
- PubMed: 27799539
- DOI: https://doi.org/10.1073/pnas.1602531113
- Primary Citation of Related Structures:
5B34, 5B35 - PubMed Abstract:
The 3D structure determination of biological macromolecules by X-ray crystallography suffers from a phase problem: to perform Fourier transformation to calculate real space density maps, both intensities and phases of structure factors are necessary; however, measured diffraction patterns give only intensities. Although serial femtosecond crystallography (SFX) using X-ray free electron lasers (XFELs) has been steadily developed since 2009, experimental phasing still remains challenging. Here, using 7.0-keV (1.771 Å) X-ray pulses from the SPring-8 Angstrom Compact Free Electron Laser (SACLA), iodine single-wavelength anomalous diffraction (SAD), single isomorphous replacement (SIR), and single isomorphous replacement with anomalous scattering (SIRAS) phasing were performed in an SFX regime for a model membrane protein bacteriorhodopsin (bR). The crystals grown in bicelles were derivatized with an iodine-labeled detergent heavy-atom additive 13a (HAD13a), which contains the magic triangle, I3C head group with three iodine atoms. The alkyl tail was essential for binding of the detergent to the surface of bR. Strong anomalous and isomorphous difference signals from HAD13a enabled successful phasing using reflections up to 2.1-Å resolution from only 3,000 and 4,000 indexed images from native and derivative crystals, respectively. When more images were merged, structure solution was possible with data truncated at 3.3-Å resolution, which is the lowest resolution among the reported cases of SFX phasing. Moreover, preliminary SFX experiment showed that HAD13a successfully derivatized the G protein-coupled A2a adenosine receptor crystallized in lipidic cubic phases. These results pave the way for de novo structure determination of membrane proteins, which often diffract poorly, even with the brightest XFEL beams.
Organizational Affiliation:
Department of Biological Sciences, Graduate School of Science, The University of Tokyo, Tokyo 113-0033, Japan.