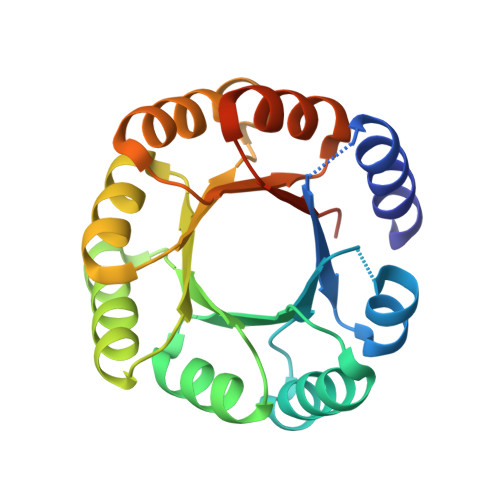
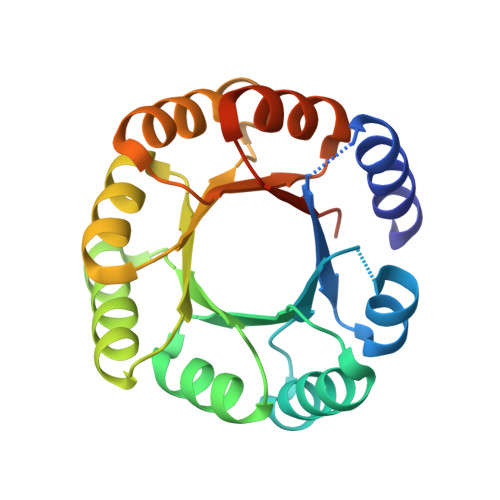
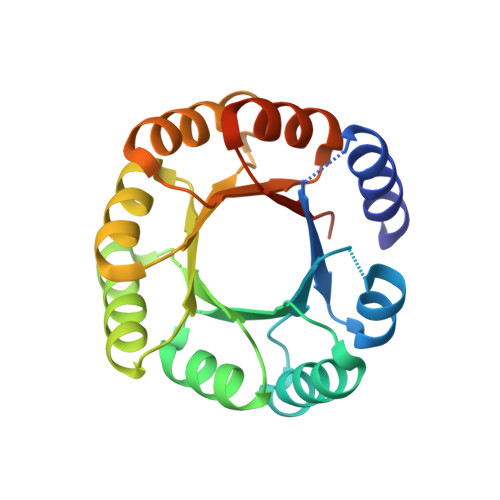
De novo design of a four-fold symmetric TIM-barrel protein with atomic-level accuracy.
Huang, P.S., Feldmeier, K., Parmeggiani, F., Fernandez Velasco, D.A., Hocker, B., Baker, D.(2016) Nat Chem Biol 12: 29-34
- PubMed: 26595462
- DOI: https://doi.org/10.1038/nchembio.1966
- Primary Citation of Related Structures:
5BVL - PubMed Abstract:
Despite efforts for over 25 years, de novo protein design has not succeeded in achieving the TIM-barrel fold. Here we describe the computational design of four-fold symmetrical (β/α)8 barrels guided by geometrical and chemical principles. Experimental characterization of 33 designs revealed the importance of side chain-backbone hydrogen bonds for defining the strand register between repeat units. The X-ray crystal structure of a designed thermostable 184-residue protein is nearly identical to that of the designed TIM-barrel model. PSI-BLAST searches do not identify sequence similarities to known TIM-barrel proteins, and sensitive profile-profile searches indicate that the design sequence is distant from other naturally occurring TIM-barrel superfamilies, suggesting that Nature has sampled only a subset of the sequence space available to the TIM-barrel fold. The ability to design TIM barrels de novo opens new possibilities for custom-made enzymes.
Organizational Affiliation:
Department of Biochemistry, University of Washington, Seattle, Washington 98195, USA.