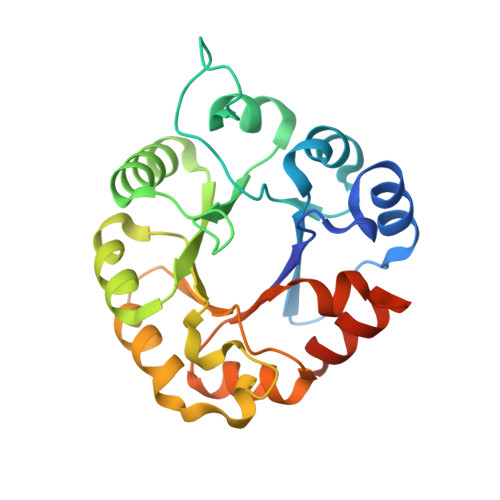
Structure and Stability of the Dimeric Triosephosphate Isomerase from the Thermophilic Archaeon Thermoplasma acidophilum.
Park, S.H., Kim, H.S., Park, M.S., Moon, S., Song, M.K., Park, H.S., Hahn, H., Kim, S.J., Bae, E., Kim, H.J., Han, B.W.(2015) PLoS One 10: e0145331-e0145331
- PubMed: 26709515
- DOI: https://doi.org/10.1371/journal.pone.0145331
- Primary Citation of Related Structures:
5CSR, 5CSS - PubMed Abstract:
Thermoplasma acidophilum is a thermophilic archaeon that uses both non-phosphorylative Entner-Doudoroff (ED) pathway and Embden-Meyerhof-Parnas (EMP) pathway for glucose degradation. While triosephosphate isomerase (TPI), a well-known glycolytic enzyme, is not involved in the ED pathway in T. acidophilum, it has been considered to play an important role in the EMP pathway. Here, we report crystal structures of apo- and glycerol-3-phosphate-bound TPI from T. acidophilum (TaTPI). TaTPI adopts the canonical TIM-barrel fold with eight α-helices and parallel eight β-strands. Although TaTPI shares ~30% sequence identity to other TPIs from thermophilic species that adopt tetrameric conformation for enzymatic activity in their harsh physiological environments, TaTPI exists as a dimer in solution. We confirmed the dimeric conformation of TaTPI by analytical ultracentrifugation and size-exclusion chromatography. Helix 5 as well as helix 4 of thermostable tetrameric TPIs have been known to play crucial roles in oligomerization, forming a hydrophobic interface. However, TaTPI contains unique charged-amino acid residues in the helix 5 and adopts dimer conformation. TaTPI exhibits the apparent Td value of 74.6°C and maintains its overall structure with some changes in the secondary structure contents at extremely acidic conditions (pH 1-2). Based on our structural and biophysical analyses of TaTPI, more compact structure of the protomer with reduced length of loops and certain patches on the surface could account for the robust nature of Thermoplasma acidophilum TPI.
Organizational Affiliation:
Research Institute of Pharmaceutical Sciences, College of Pharmacy, Seoul National University, Seoul, Korea.