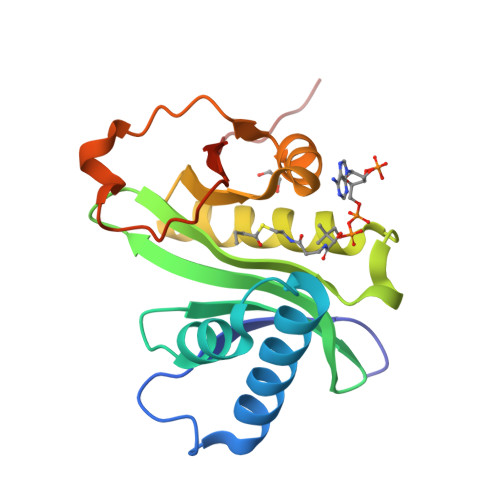
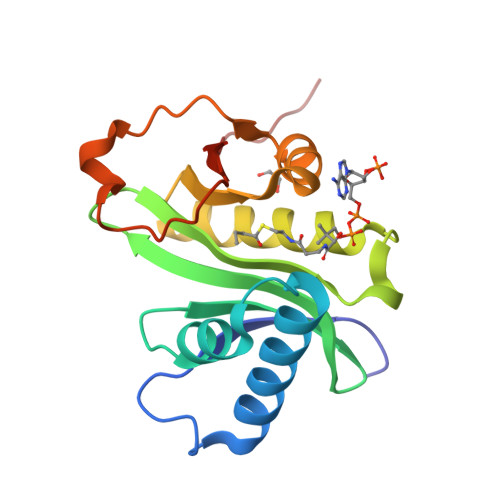
Structural basis for acyl-group discrimination by human Gcn5L2.
Ringel, A.E., Wolberger, C.(2016) Acta Crystallogr D Struct Biol 72: 841-848
- PubMed: 27377381
- DOI: https://doi.org/10.1107/S2059798316007907
- Primary Citation of Related Structures:
5H84, 5H86 - PubMed Abstract:
Gcn5 is a conserved acetyltransferase that regulates transcription by acetylating the N-terminal tails of histones. Motivated by recent studies identifying a chemically diverse array of lysine acyl modifications in vivo, the acyl-chain specificity of the acetyltransferase human Gcn5 (Gcn5L2) was examined. Whereas Gcn5L2 robustly catalyzes lysine acetylation, the acyltransferase activity of Gcn5L2 becomes progressively weaker with increasing acyl-chain length. To understand how Gcn5 discriminates between different acyl-CoA molecules, structures of the catalytic domain of human Gcn5L2 bound to propionyl-CoA and butyryl-CoA were determined. Although the active site of Gcn5L2 can accommodate propionyl-CoA and butyryl-CoA without major structural rearrangements, butyryl-CoA adopts a conformation incompatible with catalysis that obstructs the path of the incoming lysine residue and acts as a competitive inhibitor of Gcn5L2 versus acetyl-CoA. These structures demonstrate how Gcn5L2 discriminates between acyl-chain donors and explain why Gcn5L2 has weak activity for acyl moieties that are larger than an acetyl group.
Organizational Affiliation:
Department of Biophysics and Biophysical Chemistry, Johns Hopkins University School of Medicine, 725 North Wolfe Street, Baltimore, MD 21205, USA.