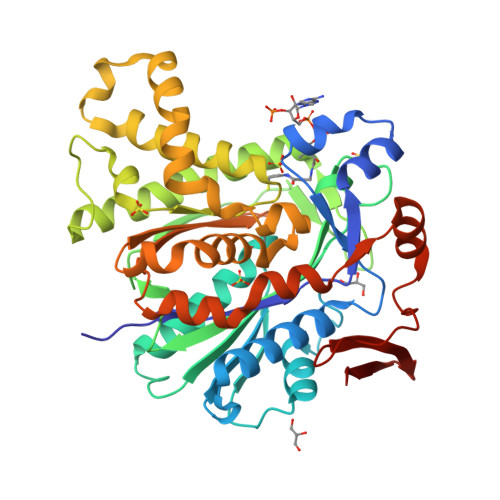
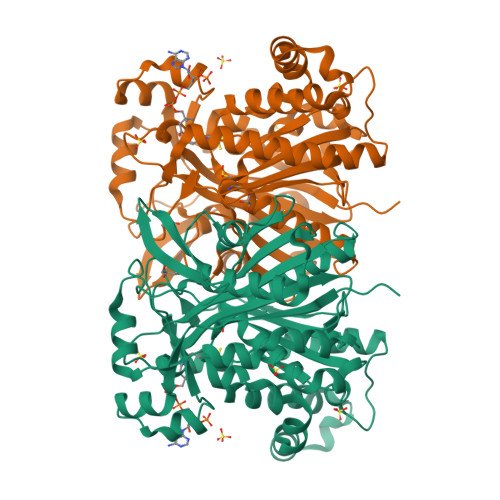
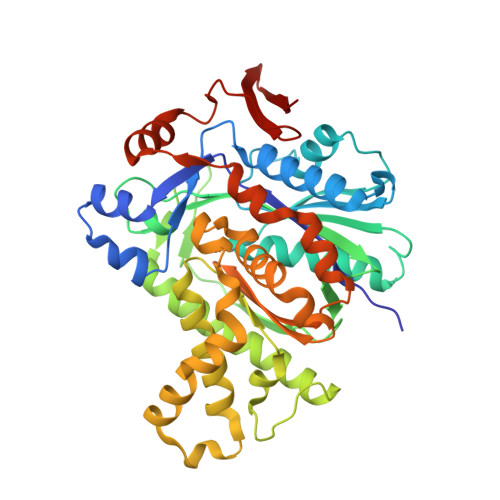
Crystal Structure of the HMG-CoA Synthase MvaS from the Gram-Negative Bacterium Myxococcus xanthus.
Bock, T., Kasten, J., Muller, R., Blankenfeldt, W.(2016) Chembiochem 17: 1257-1262
- PubMed: 27124816
- DOI: https://doi.org/10.1002/cbic.201600070
- Primary Citation of Related Structures:
5HWO, 5HWP, 5HWQ, 5HWR - PubMed Abstract:
A critical step in bacterial isoprenoid production is the synthesis of 3-hydroxy-3-methylglutaryl coenzyme A (HMG-CoA) catalyzed by HMG-CoA synthase (HMGCS). In myxobacteria, this enzyme is also involved in a recently discovered alternative and acetyl-CoA-dependent isovaleryl CoA biosynthesis pathway. Here we present crystal structures of MvaS, the HMGCS from Myxococcus xanthus, in complex with CoA and acetylated active site Cys115, with the second substrate acetoacetyl CoA and with the product of the condensation reaction, 3-hydroxy-3-methylglutaryl CoA. With these structures, we show that MvaS uses the common HMGCS enzymatic mechanism and provide evidence that dimerization plays a role in the formation and stability of the active site. Overall, MvaS shows features typical of the eukaryotic HMGCS and exhibits differences from homologues from Gram-positive bacteria. This study provides insights into myxobacterial alternative isovaleryl CoA biosynthesis and thereby extends the toolbox for the biotechnological production of renewable fuel and chemicals.
Organizational Affiliation:
Structure and Function of Proteins, Helmholtz Centre for Infection Research, Inhoffenstrasse 7, 38124, Braunschweig, Germany.