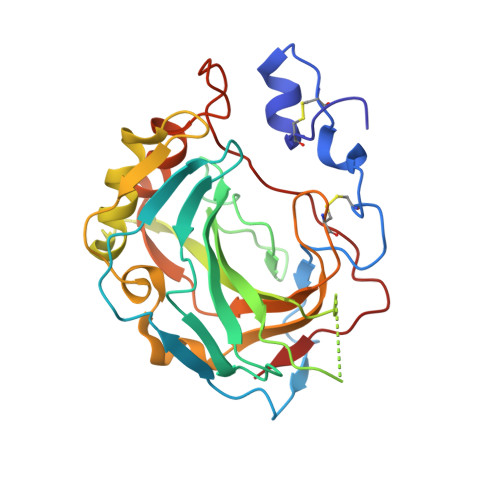
Intrinsic thermodynamics of high affinity inhibitor binding to recombinant human carbonic anhydrase IV.
Mickeviciute, A., Timm, D.D., Gedgaudas, M., Linkuviene, V., Chen, Z., Waheed, A., Michailoviene, V., Zubriene, A., Smirnov, A., Capkauskaite, E., Baranauskiene, L., Jachno, J., Revuckiene, J., Manakova, E., Grazulis, S., Matuliene, J., Di Cera, E., Sly, W.S., Matulis, D.(2018) Eur Biophys J 47: 271-290
- PubMed: 28975383
- DOI: https://doi.org/10.1007/s00249-017-1256-0
- Primary Citation of Related Structures:
5IPZ, 5JN8, 5JN9, 5JNA, 5JNC, 5KU6 - PubMed Abstract:
Membrane-associated carbonic anhydrase (CA) isoform IV participates in carbon metabolism and pH homeostasis and is implicated in the development of eye diseases such as retinitis pigmentosa and glaucoma. A series of substituted benzenesulfonamides were designed and their binding affinity to CA IV was determined by fluorescent thermal shift assay and isothermal titration calorimetry (ITC). Compound [(4-chloro-2-phenylsulfanyl-5-sulfamoyl-benzoyl)amino]propyl acetate (19) bound CA IV with the K d of 1.0 nM and exhibited significant selectivity over the remaining 11 human CA isoforms. The compound could be developed as a drug targeting CA IV. Various forms of recombinant CA IV were produced in Escherichia coli and mammalian cell cultures. Comparison of their temperature stability in various buffers and salt solutions demonstrated that CA IV is most stable at slightly alkaline conditions and at elevated sodium sulfate concentrations. High-resolution X-ray crystallographic structures of ortho-Cl and meta-thiazole-substituted benzene sulfonamide in complex with CA IV revealed the position of and interactions between the ligand and the protein. Sulfonamide inhibitor binding to CA IV is linked to several reactions-the deprotonation of the sulfonamide amino group, the protonation of CA-Zn(II)-bound hydroxide at the active site of CA IV, and the compensating reactions of the buffer. The dissection of binding-linked reactions yielded the intrinsic thermodynamic parameters, characterizing the interaction between CA IV and the sulfonamides in the binding-able protonation forms, including Gibbs energy, enthalpy, and entropy, that could be used for the characterization of binding to any CA in the process of drug design.
Organizational Affiliation:
Department of Biothermodynamics and Drug Design, Institute of Biotechnology, Life Sciences Center, Vilnius University, Saulėtekio 7, 10257, Vilnius, Lithuania.