Catalase-peroxidase KatG of Burkholderia pseudomallei at 1.7A resolution.
Carpena, X., Loprasert, S., Mongkolsuk, S., Switala, J., Loewen, P.C., Fita, I.(2003) J Mol Biol 327: 475-489
- PubMed: 12628252
- DOI: https://doi.org/10.1016/s0022-2836(03)00122-0
- Primary Citation of Related Structures:
5L05 - PubMed Abstract:
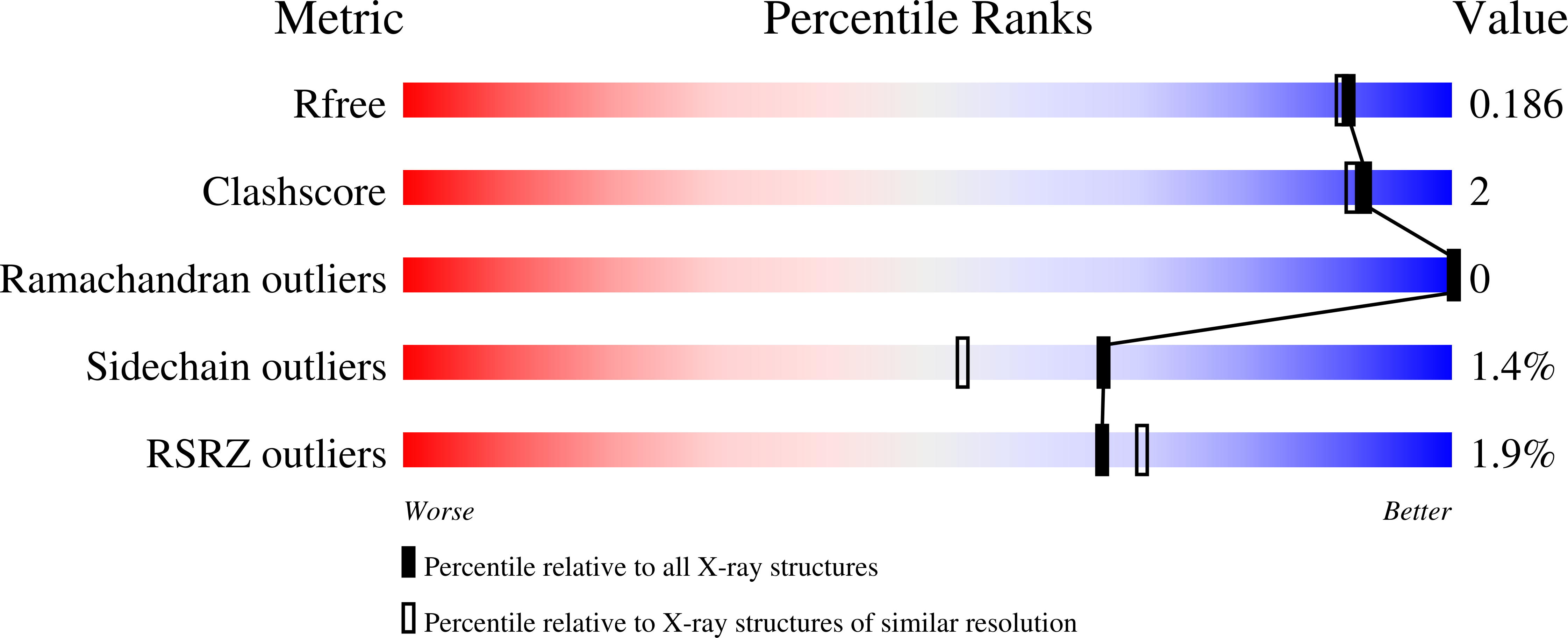
The catalase-peroxidase encoded by katG of Burkholderia pseudomallei (BpKatG) is 65% identical with KatG of Mycobacterium tuberculosis, the enzyme responsible for the activation of isoniazid as an antibiotic. The structure of a complex of BpKatG with an unidentified ligand, has been solved and refined at 1.7A resolution using X-ray synchrotron data collected from crystals flash-cooled with liquid nitrogen. The crystallographic agreement factors R and R(free) are 15.3% and 18.6%, respectively. The crystallized enzyme is a dimer with one modified heme group and one metal ion, likely sodium, per subunit. The modification on the heme group involves the covalent addition of two or three atoms, likely a perhydroxy group, to the secondary carbon atom of the vinyl group on ring I. The added group can form hydrogen bonds with two water molecules that are also in contact with the active-site residues Trp111 and His112, suggesting that the modification may have a catalytic role. The heme modification is in close proximity to an unusual covalent adduct among the side-chains of Trp111, Tyr238 and Met264. In addition, Trp111 appears to be oxidized on C(delta1) of the indole ring. The main channel, providing access of substrate hydrogen peroxide to the heme, contains a region of unassigned electron density consistent with the binding of a pyridine nucleotide-like molecule. An interior cavity, containing the sodium ion and an additional region of unassigned density, is evident adjacent to the adduct and is accessible to the outside through a second funnel-shaped channel. A large cleft in the side of the subunit is evident and may be a potential substrate-binding site with a clear pathway for electron transfer to the active-site heme group through the adduct.
Organizational Affiliation:
CID-CSIC, Jordi-Girona 18-26, 08034 Barcelona, Spain.