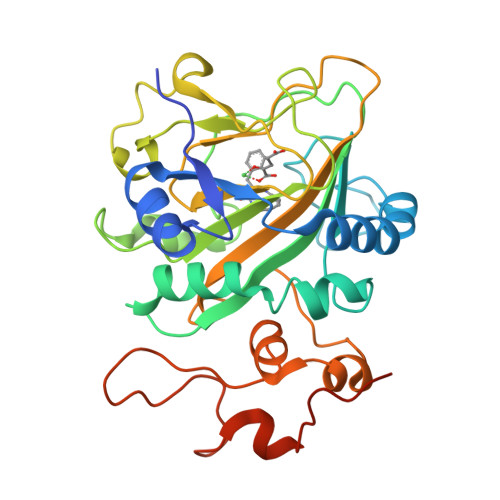
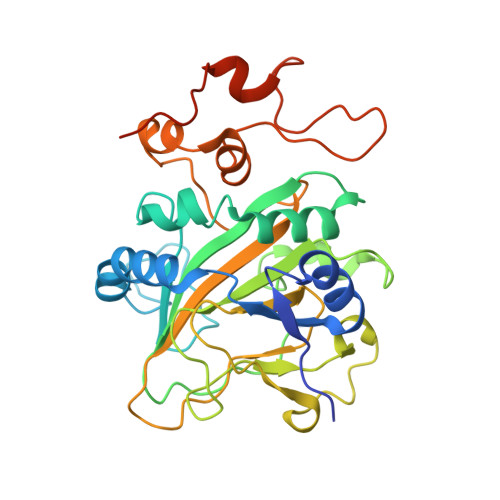
Catalytic mechanism and molecular engineering of quinolone biosynthesis in dioxygenase AsqJ.
Mader, S.L., Brauer, A., Groll, M., Kaila, V.R.I.(2018) Nat Commun 9: 1168-1168
- PubMed: 29563492
- DOI: https://doi.org/10.1038/s41467-018-03442-2
- Primary Citation of Related Structures:
5OA4, 5OA7, 5OA8, 6EOZ - PubMed Abstract:
The recently discovered Fe II /α-ketoglutarate-dependent dioxygenase AsqJ from Aspergillus nidulans stereoselectively catalyzes a multistep synthesis of quinolone alkaloids, natural products with significant biomedical applications. To probe molecular mechanisms of this elusive catalytic process, we combine here multi-scale quantum and classical molecular simulations with X-ray crystallography, and in vitro biochemical activity studies. We discover that methylation of the substrate is essential for the activity of AsqJ, establishing molecular strain that fine-tunes π-stacking interactions within the active site. To rationally engineer AsqJ for modified substrates, we amplify dispersive interactions within the active site. We demonstrate that the engineered enzyme has a drastically enhanced catalytic activity for non-methylated surrogates, confirming our computational data and resolved high-resolution X-ray structures at 1.55 Å resolution. Our combined findings provide crucial mechanistic understanding of the function of AsqJ and showcase how combination of computational and experimental data enables to rationally engineer enzymes.
Organizational Affiliation:
Center for Integrated Protein Science Munich (CIPSM), Department Chemie, Technische Universität München, Lichtenbergstraße 4, 85748, Garching, Germany.