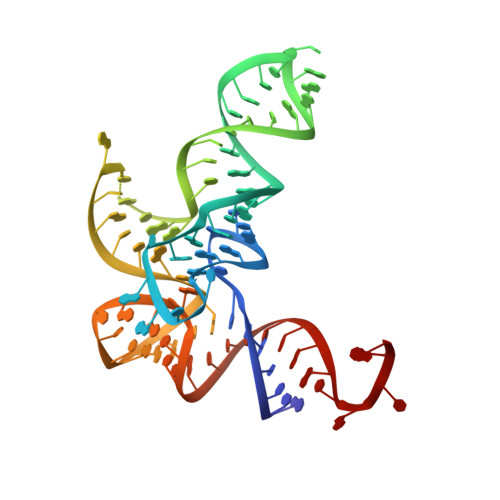
Kinetic Origin of Substrate Specificity in Post-Transfer Editing by Leucyl-tRNA Synthetase.
Dulic, M., Cvetesic, N., Zivkovic, I., Palencia, A., Cusack, S., Bertosa, B., Gruic-Sovulj, I.(2018) J Mol Biol 430: 1-16
- PubMed: 29111343
- DOI: https://doi.org/10.1016/j.jmb.2017.10.024
- Primary Citation of Related Structures:
5OMW, 5ON2, 5ON3, 5ONH - PubMed Abstract:
The intrinsic editing capacities of aminoacyl-tRNA synthetases ensure a high-fidelity translation of the amino acids that possess effective non-cognate aminoacylation surrogates. The dominant error-correction pathway comprises deacylation of misaminoacylated tRNA within the aminoacyl-tRNA synthetase editing site. To assess the origin of specificity of Escherichia coli leucyl-tRNA synthetase (LeuRS) against the cognate aminoacylation product in editing, we followed binding and catalysis independently using cognate leucyl- and non-cognate norvalyl-tRNA Leu and their non-hydrolyzable analogues. We found that the amino acid part (leucine versus norvaline) of (mis)aminoacyl-tRNAs can contribute approximately 10-fold to ground-state discrimination at the editing site. In sharp contrast, the rate of deacylation of leucyl- and norvalyl-tRNA Leu differed by about 10 4 -fold. We further established the critical role for the A76 3'-OH group of the tRNA Leu in post-transfer editing, which supports the substrate-assisted deacylation mechanism. Interestingly, the abrogation of the LeuRS specificity determinant threonine 252 did not improve the affinity of the editing site for the cognate leucine as expected, but instead substantially enhanced the rate of leucyl-tRNA Leu hydrolysis. In line with that, molecular dynamics simulations revealed that the wild-type enzyme, but not the T252A mutant, enforced leucine to adopt the side-chain conformation that promotes the steric exclusion of a putative catalytic water. Our data demonstrated that the LeuRS editing site exhibits amino acid specificity of kinetic origin, arguing against the anticipated prominent role of steric exclusion in the rejection of leucine. This feature distinguishes editing from the synthetic site, which relies on ground-state discrimination in amino acid selection.
Organizational Affiliation:
Department of Chemistry, Faculty of Science, University of Zagreb, Horvatovac 102a, 10000 Zagreb, Croatia.