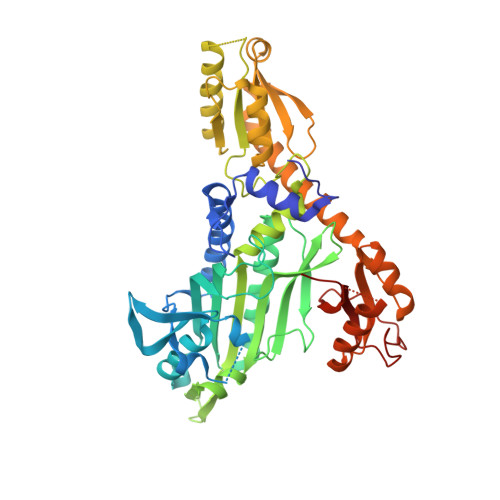
Targeting Prolyl-tRNA Synthetase to Accelerate Drug Discovery against Malaria, Leishmaniasis, Toxoplasmosis, Cryptosporidiosis, and Coccidiosis
Jain, V., Yogavel, M., Kikuchi, H., Oshima, Y., Hariguchi, N., Matsumoto, M., Goel, P., Touquet, B., Jumani, R.S., Tacchini-Cottier, F., Harlos, K., Huston, C.D., Hakimi, M.A., Sharma, A.(2017) Structure 25: 1495-1505.e6
- PubMed: 28867614
- DOI: https://doi.org/10.1016/j.str.2017.07.015
- Primary Citation of Related Structures:
5XIF, 5XIG, 5XIH, 5XII, 5XIJ, 5XIK, 5XIL, 5XIO, 5XIP, 5XIQ - PubMed Abstract:
Developing anti-parasitic lead compounds that act on key vulnerabilities are necessary for new anti-infectives. Malaria, leishmaniasis, toxoplasmosis, cryptosporidiosis and coccidiosis together kill >500,000 humans annually. Their causative parasites Plasmodium, Leishmania, Toxoplasma, Cryptosporidium and Eimeria display high conservation in many housekeeping genes, suggesting that these parasites can be attacked by targeting invariant essential proteins. Here, we describe selective and potent inhibition of prolyl-tRNA synthetases (PRSs) from the above parasites using a series of quinazolinone-scaffold compounds. Our PRS-drug co-crystal structures reveal remarkable active site plasticity that accommodates diversely substituted compounds, an enzymatic feature that can be leveraged for refining drug-like properties of quinazolinones on a per parasite basis. A compound we termed In-5 exhibited a unique double conformation, enhanced drug-like properties, and cleared malaria in mice. It thus represents a new lead for optimization. Collectively, our data offer insights into the structure-guided optimization of quinazolinone-based compounds for drug development against multiple human eukaryotic pathogens.
Organizational Affiliation:
Molecular Medicine - Structural Parasitology Group, International Centre for Genetic Engineering and Biotechnology (ICGEB), Aruna Asaf Ali Marg, New Delhi 110067, India.