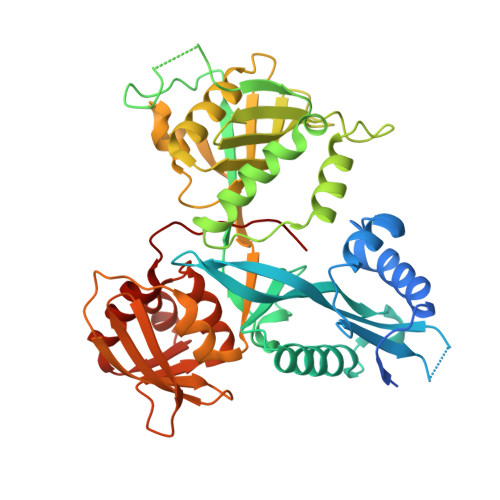
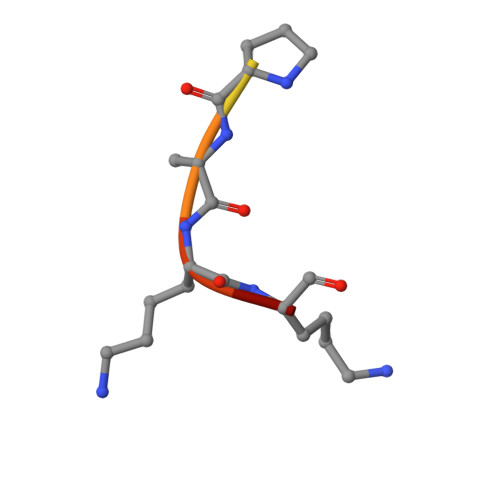
Acetylation by Eis and Deacetylation by Rv1151c of Mycobacterium tuberculosis HupB: Biochemical and Structural Insight.
Green, K.D., Biswas, T., Pang, A.H., Willby, M.J., Reed, M.S., Stuchlik, O., Pohl, J., Posey, J.E., Tsodikov, O.V., Garneau-Tsodikova, S.(2018) Biochemistry 57: 781-790
- PubMed: 29345920
- DOI: https://doi.org/10.1021/acs.biochem.7b01089
- Primary Citation of Related Structures:
6B0U - PubMed Abstract:
Bacterial nucleoid-associated proteins (NAPs) are critical to genome integrity and chromosome maintenance. Post-translational modifications of bacterial NAPs appear to function similarly to their better studied mammalian counterparts. The histone-like NAP HupB from Mycobacterium tuberculosis (Mtb) was previously observed to be acetylated by the acetyltransferase Eis, leading to genome reorganization. We report biochemical and structural aspects of acetylation of HupB by Eis. We also found that the SirT-family NAD + -dependent deacetylase Rv1151c from Mtb deacetylated HupB in vitro and characterized the deacetylation kinetics. We propose that activities of Eis and Rv1151c could regulate the acetylation status of HupB to remodel the mycobacterial chromosome in response to environmental changes.
Organizational Affiliation:
Department of Pharmaceutical Sciences, College of Pharmacy, University of Kentucky , Lexington, Kentucky 40536-0596, United States.