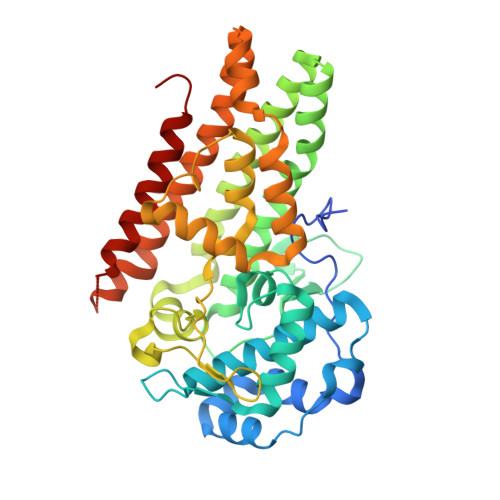
High-resolution structures of inhibitor complexes of human indoleamine 2,3-dioxygenase 1 in a new crystal form.
Luo, S., Xu, K., Xiang, S., Chen, J., Chen, C., Guo, C., Tong, Y., Tong, L.(2018) Acta Crystallogr F Struct Biol Commun 74: 717-724
- PubMed: 30387777
- DOI: https://doi.org/10.1107/S2053230X18012955
- Primary Citation of Related Structures:
6E40, 6E41, 6E42, 6E43, 6E44, 6E45, 6E46 - PubMed Abstract:
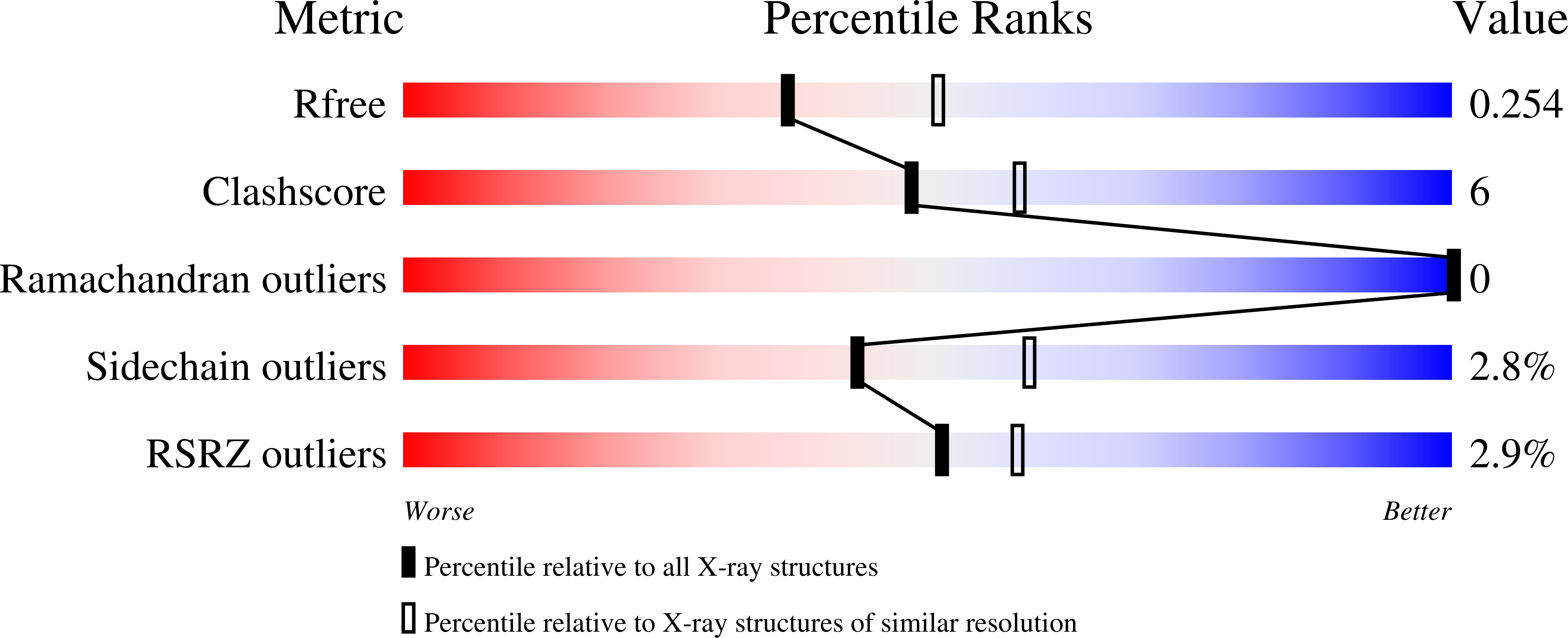
Human indoleamine 2,3-dioxygenase 1 (IDO1) is a heme-dependent enzyme with important roles in many cellular processes and is a potential target for drug discovery against cancer and other diseases. Crystal structures of IDO1 in complex with various inhibitors have been reported. Many of these crystals belong to the same crystal form and most of the reported structures have resolutions in the range 3.2-2.3 Å. Here, three new crystal forms of human IDO1 obtained by introducing a surface mutation, K116A/K117A, distant from the active site are reported. One of these crystal forms diffracted to 1.5 Å resolution and can be readily used for soaking experiments to determine high-resolution structures of IDO1 in complex with the substrate tryptophan or inhibitors that coordinate the heme. In addition, this mutant was used to produce crystals of a complex with an inhibitor that targets the apo form of the enzyme under the same conditions; the structure of this complex was determined at 1.7 Å resolution. Overall, this mutant represents a robust platform for determining the structures of inhibitor and substrate complexes of IDO1 at high resolution.
Organizational Affiliation:
Department of Biological Sciences, Columbia University, New York, NY 10027, USA.