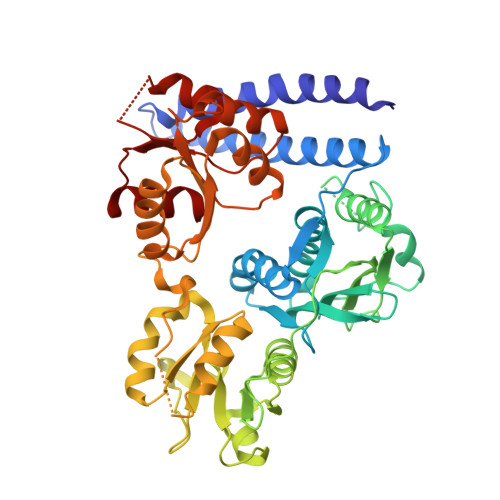
Hybrid histidine kinase activation by cyclic di-GMP-mediated domain liberation.
Dubey, B.N., Agustoni, E., Bohm, R., Kaczmarczyk, A., Mangia, F., von Arx, C., Jenal, U., Hiller, S., Plaza-Menacho, I., Schirmer, T.(2020) Proc Natl Acad Sci U S A 117: 1000-1008
- PubMed: 31882446
- DOI: https://doi.org/10.1073/pnas.1911427117
- Primary Citation of Related Structures:
6QRJ, 6QRL - PubMed Abstract:
Cytosolic hybrid histidine kinases (HHKs) constitute major signaling nodes that control various biological processes, but their input signals and how these are processed are largely unknown. In Caulobacter crescentus , the HHK ShkA is essential for accurate timing of the G1-S cell cycle transition and is regulated by the corresponding increase in the level of the second messenger c-di-GMP. Here, we use a combination of X-ray crystallography, NMR spectroscopy, functional analyses, and kinetic modeling to reveal the regulatory mechanism of ShkA. In the absence of c-di-GMP, ShkA predominantly adopts a compact domain arrangement that is catalytically inactive. C-di-GMP binds to the dedicated pseudoreceiver domain Rec1, thereby liberating the canonical Rec2 domain from its central position where it obstructs the large-scale motions required for catalysis. Thus, c-di-GMP cannot only stabilize domain interactions, but also engage in domain dissociation to allosterically invoke a downstream effect. Enzyme kinetics data are consistent with conformational selection of the ensemble of active domain constellations by the ligand and show that autophosphorylation is a reversible process.
Organizational Affiliation:
Structural Biology, Biozentrum, University of Basel, 4056 Basel, Switzerland.