Discovery of novel Cyclophilin D inhibitors starting from three dimensional fragments with millimolar potencies.
Gradler, U., Schwarz, D., Blaesse, M., Leuthner, B., Johnson, T.L., Bernard, F., Jiang, X., Marx, A., Gilardone, M., Lemoine, H., Roche, D., Jorand-Lebrun, C.(2019) Bioorg Med Chem Lett 29: 126717-126717
- PubMed: 31635932
- DOI: https://doi.org/10.1016/j.bmcl.2019.126717
- Primary Citation of Related Structures:
6R8L, 6R8O, 6R8W, 6R9S, 6R9U, 6R9X, 6RA1 - PubMed Abstract:
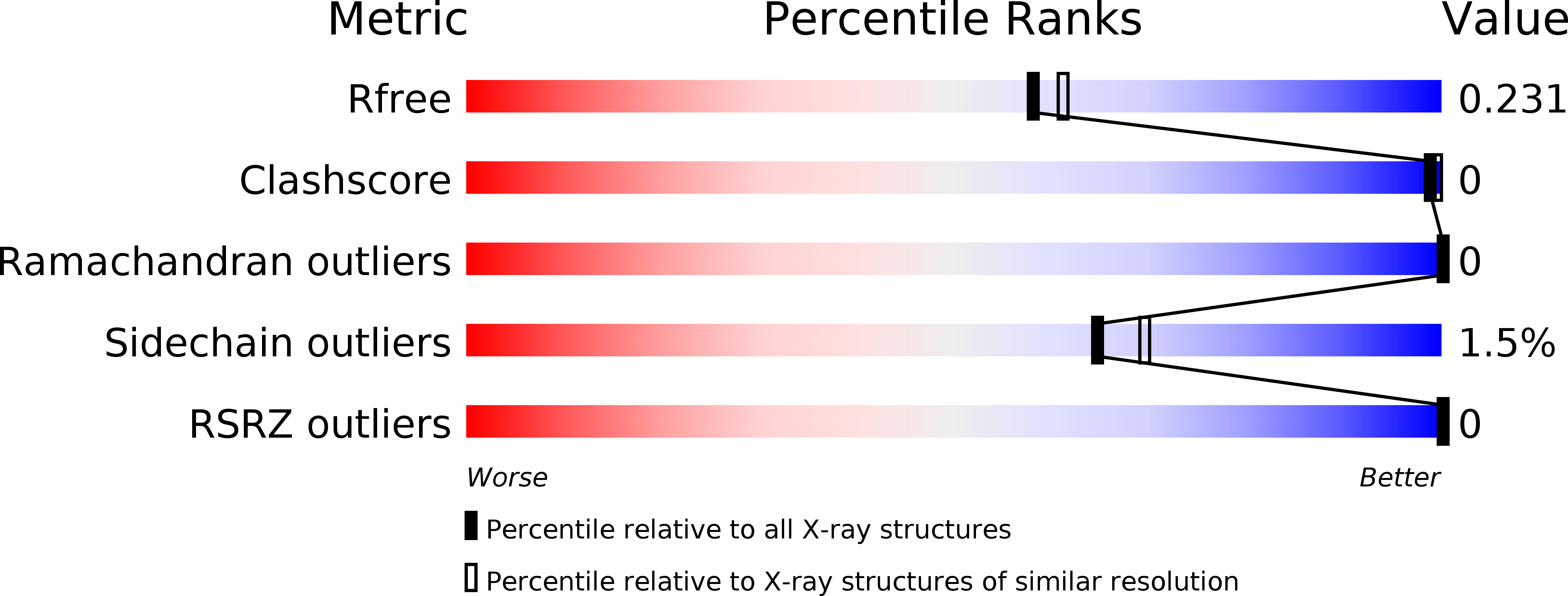
Fragment-based screening by SPR enabled the discovery of chemical diverse fragment hits with millimolar binding affinities to the peptidyl-prolyl isomerase Cyclophilin D (CypD). The CypD protein crystal structures of 6 fragment hits provided the basis for subsequent medicinal chemistry optimization by fragment merging and linking yielding three different chemical series with either urea, oxalyl or amide linkers connecting millimolar fragments in the S1' and S2 pockets. We successfully improved the in vitro CypD potencies in the biochemical FP and PPIase assays and in the biophysical SPR binding assay from millimolar towards the low micromolar and submicromolar range by >1000-fold for some fragment derivatives. The initial SAR together with the protein crystal structures of our novel CypD inhibitors provide a suitable basis for further hit-to-lead optimization.
Organizational Affiliation:
Merck Healthcare, Merck KGaA, Frankfurter Str. 250, D-64293 Darmstadt, Germany. Electronic address: ulrich.graedler@merckgroup.com.