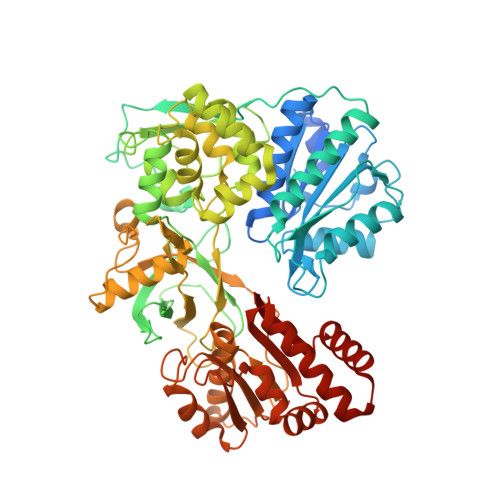
Biochemical and structural insights into the cytochrome P450 reductase from Candida tropicalis.
Ebrecht, A.C., van der Bergh, N., Harrison, S.T.L., Smit, M.S., Sewell, B.T., Opperman, D.J.(2019) Sci Rep 9: 20088-20088
- PubMed: 31882753
- DOI: https://doi.org/10.1038/s41598-019-56516-6
- Primary Citation of Related Structures:
6T1T, 6T1U - PubMed Abstract:
Cytochrome P450 reductases (CPRs) are diflavin oxidoreductases that supply electrons to type II cytochrome P450 monooxygenases (CYPs). In addition, it can also reduce other proteins and molecules, including cytochrome c, ferricyanide, and different drugs. Although various CPRs have been functionally and structurally characterized, the overall mechanism and its interaction with different redox acceptors remain elusive. One of the main problems regarding electron transfer between CPRs and CYPs is the so-called "uncoupling", whereby NAD(P)H derived electrons are lost due to the reduced intermediates' (FAD and FMN of CPR) interaction with molecular oxygen. Additionally, the decay of the iron-oxygen complex of the CYP can also contribute to loss of reducing equivalents during an unproductive reaction cycle. This phenomenon generates reactive oxygen species (ROS), leading to an inefficient reaction. Here, we present the study of the CPR from Candida tropicalis (CtCPR) lacking the hydrophobic N-terminal part (Δ2-22). The enzyme supports the reduction of cytochrome c and ferricyanide, with an estimated 30% uncoupling during the reactions with cytochrome c. The ROS produced was not influenced by different physicochemical conditions (ionic strength, pH, temperature). The X-ray structures of the enzyme were solved with and without its cofactor, NADPH. Both CtCPR structures exhibited the closed conformation. Comparison with the different solved structures revealed an intricate ionic network responsible for the regulation of the open/closed movement of CtCPR.
Organizational Affiliation:
Department of Microbial, Biochemical, and Food Biotechnology, University of the Free State, Bloemfontein, 9301, South Africa.