Cocktail experiment E: fragments 52, 58, and 63 at 90 mM concentration in complex with Endothiapepsin
Hassaan, E., Klebe, G., Heine, A., Schiebel, J., Koester, H., Softley, C., Sattler, M.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Endothiapepsin | 419 | Cryphonectria parasitica | Mutation(s): 0 EC: 3.4.23.22 | 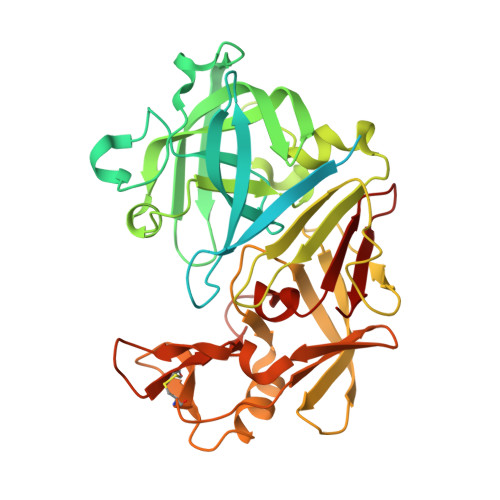 | |
UniProt | |||||
Find proteins for P11838 (Cryphonectria parasitica) Explore P11838 Go to UniProtKB: P11838 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P11838 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| F63 (Subject of Investigation/LOI) Query on F63 | B [auth A] | N-methyl-1-[5-(pyridin-3-yloxy)furan-2-yl]methanamine C11 H12 N2 O2 MUKCVLFJCGZPKF-UHFFFAOYSA-N |  | ||
| GOL Query on GOL | G [auth A] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| DMS Query on DMS | C [auth A], D [auth A], E [auth A], F [auth A] | DIMETHYL SULFOXIDE C2 H6 O S IAZDPXIOMUYVGZ-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 45.401 | α = 90 |
| b = 73.178 | β = 109.762 |
| c = 52.659 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| Coot | model building |
| XSCALE | data scaling |
| PHASER | phasing |
| XDS | data reduction |