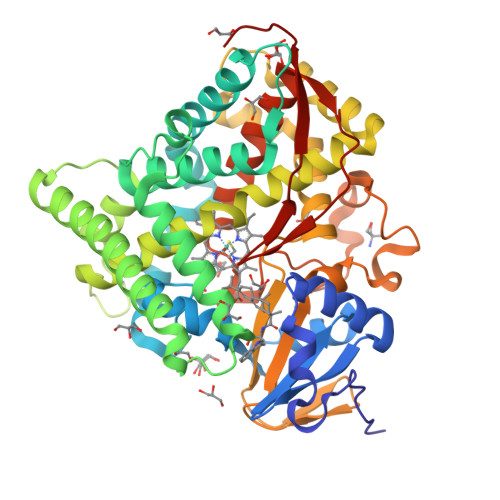
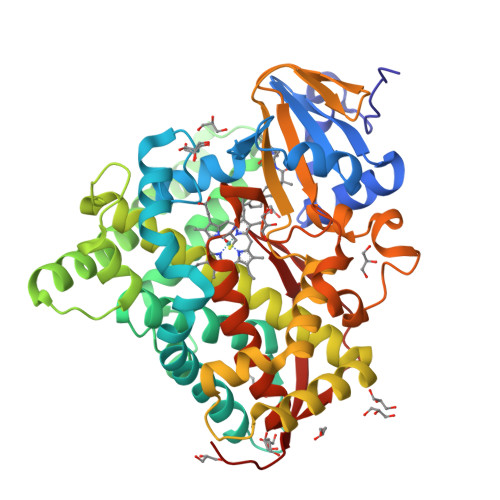
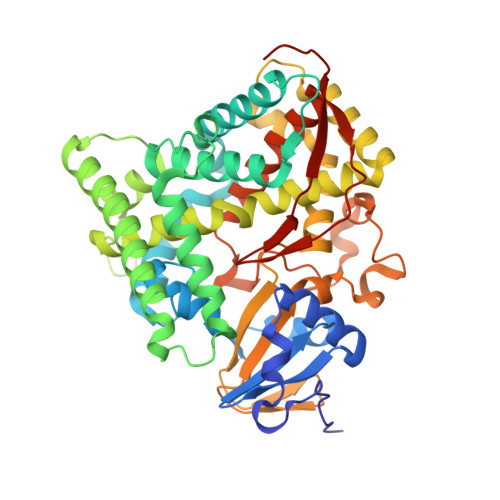
Crystals in Minutes: Instant On-Site Microcrystallisation of Various Flavours of the CYP102A1 (P450BM3) Haem Domain.
Stanfield, J.K., Omura, K., Matsumoto, A., Kasai, C., Sugimoto, H., Shiro, Y., Watanabe, Y., Shoji, O.(2020) Angew Chem Int Ed Engl 59: 7611-7618
- PubMed: 32157795
- DOI: https://doi.org/10.1002/anie.201913407
- Primary Citation of Related Structures:
6JLV, 6JMH, 6JMW, 6JO1, 6JS8, 6JVC, 6JZS, 6K24, 6K58, 6K9S - PubMed Abstract:
Despite CYP102A1 (P450BM3) representing one of the most extensively researched metalloenzymes, crystallisation of its haem domain upon modification can be a challenge. Crystal structures are indispensable for the efficient structure-based design of P450BM3 as a biocatalyst. The abietane diterpenoid derivative N-abietoyl-l-tryptophan (AbiATrp) is an outstanding crystallisation accelerator for the wild-type P450BM3 haem domain, with visible crystals forming within 2 hours and diffracting to a near-atomic resolution of 1.22 Å. Using these crystals as seeds in a cross-microseeding approach, an assortment of P450BM3 haem domain crystal structures, containing previously uncrystallisable decoy molecules and diverse artificial metalloporphyrins binding various ligand molecules, as well as heavily tagged haem-domain variants, could be determined. Some of the structures reported herein could be used as models of different stages of the P450BM3 catalytic cycle.
Organizational Affiliation:
Department of Chemistry, Graduate School of Science, Nagoya University, Furo-cho, Chikusa-ku, Nagoya, 464-8602, Japan.