Ultrapotent neutralizing antibodies against SARS-CoV-2 with a high degree of mutation resistance.
Zou, J., Li, L., Zheng, P., Liang, W., Hu, S., Zhou, S., Wang, Y., Zhao, J., Yuan, D., Liu, L., Wu, D., Xu, M., Zhang, F., Zhu, M., Wu, Z., Cao, X., Ni, M., Ling, X., Wu, Y., Kuang, Z., Hu, M., Li, J., Li, X., Guo, X., Xu, T., Jiang, H., Gao, C., Yu, M., Liu, J., Zhong, N., Zhou, J., Huang, J.A., Jin, T., He, J.(2022) J Clin Invest 132
- PubMed: 35108220
- DOI: https://doi.org/10.1172/JCI154987
- Primary Citation of Related Structures:
7FCP, 7FCQ - PubMed Abstract:
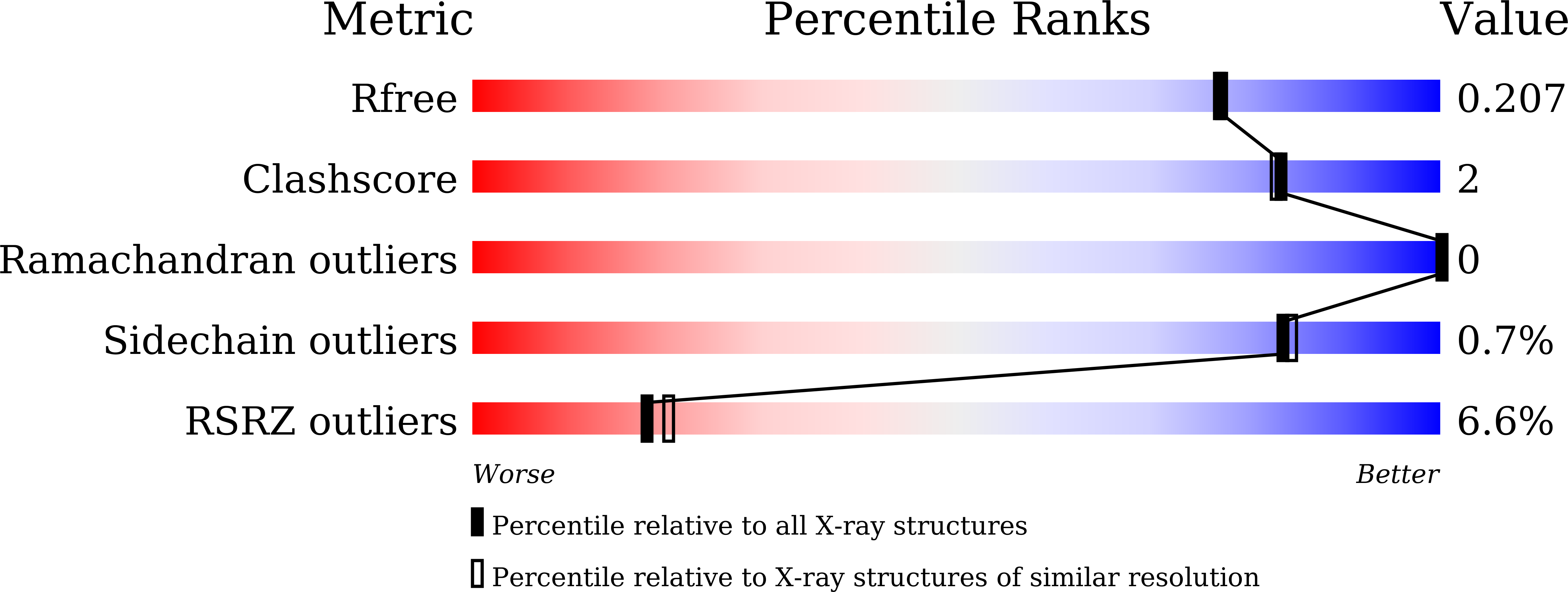
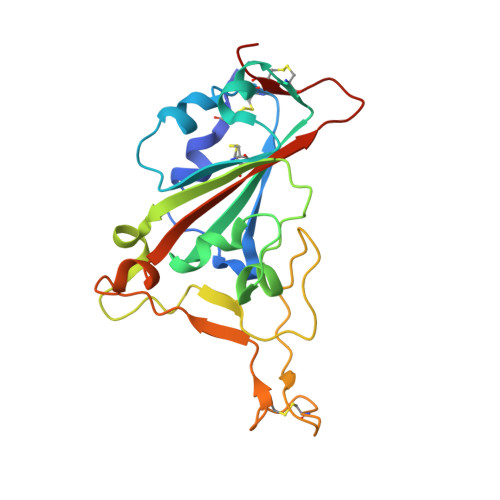
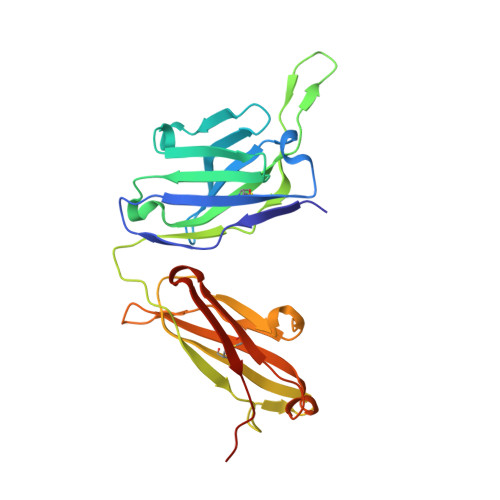
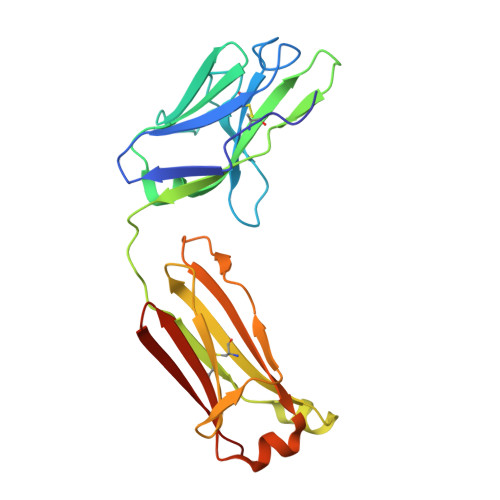
Many SARS-CoV-2 neutralizing antibodies (nAbs) lose potency against variants of concern. In this study, we developed 2 strategies to produce mutation-resistant antibodies. First, a yeast library expressing mutant receptor binding domains (RBDs) of the spike protein was utilized to screen for potent nAbs that are least susceptible to viral escape. Among the candidate antibodies, P5-22 displayed ultrahigh potency for virus neutralization as well as an outstanding mutation resistance profile. Additionally, P14-44 and P15-16 were recognized as mutation-resistant antibodies with broad betacoronavirus neutralization properties. P15-16 has only 1 binding hotspot, which is K378 in the RBD of SARS-CoV-2. The crystal structure of the P5-22, P14-44, and RBD ternary complex clarified the unique mechanisms that underlie the excellent mutation resistance profiles of these antibodies. Secondly, polymeric IgG enhanced antibody avidity by eliminating P5-22's only hotspot, residue F486 in the RBD, thereby potently blocking cell entry by mutant viruses. Structural and functional analyses of antibodies screened using both potency assays and the yeast RBD library revealed rare, ultrapotent, mutation-resistant nAbs against SARS-CoV-2.
Organizational Affiliation:
Guangzhou Laboratory, Guangzhou International Biotech Island, Guangzhou, China.