Rational Design of Hybrid SARS-CoV-2 Main Protease Inhibitors Guided by the Superimposed Cocrystal Structures with the Peptidomimetic Inhibitors GC-376, Telaprevir, and Boceprevir.
Xia, Z., Sacco, M., Hu, Y., Ma, C., Meng, X., Zhang, F., Szeto, T., Xiang, Y., Chen, Y., Wang, J.(2021) Acs Pharmacol Transl Sci 4: 1408-1421
- PubMed: 34414360
- DOI: https://doi.org/10.1021/acsptsci.1c00099
- Primary Citation of Related Structures:
7LYH, 7LYI - PubMed Abstract:
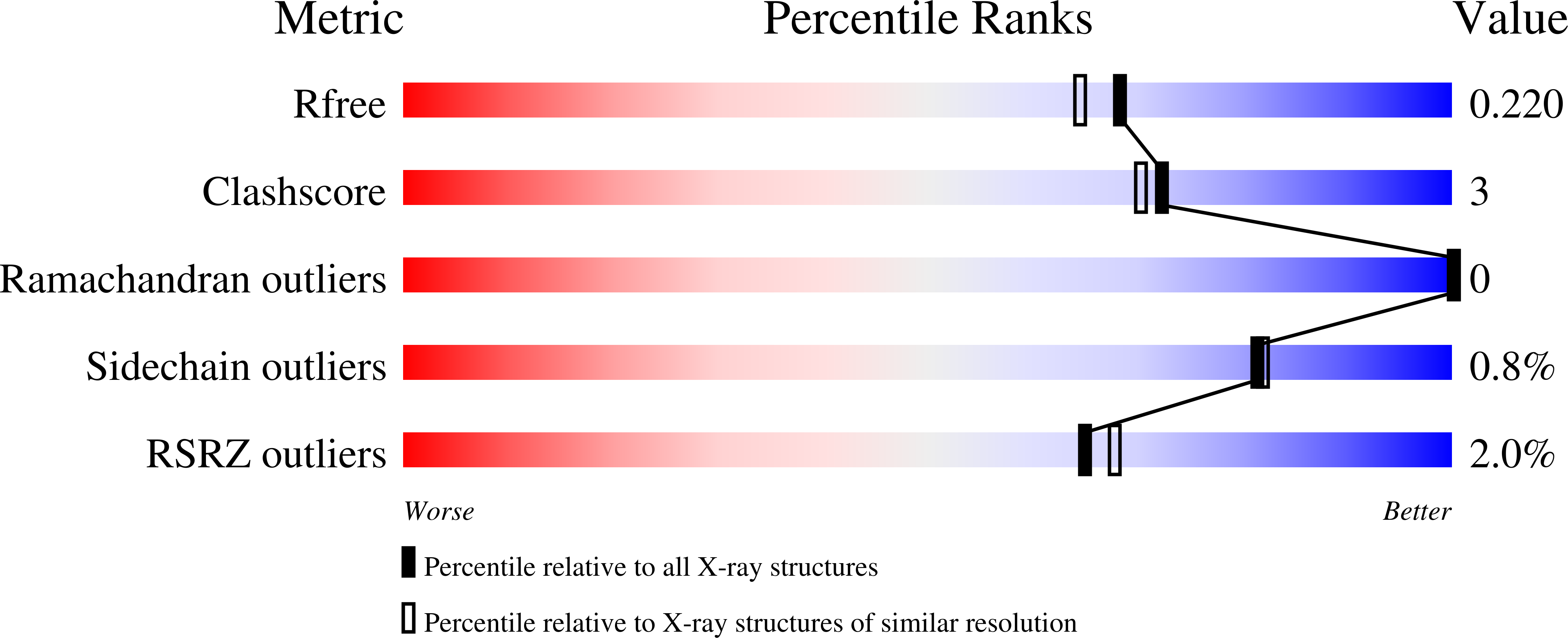
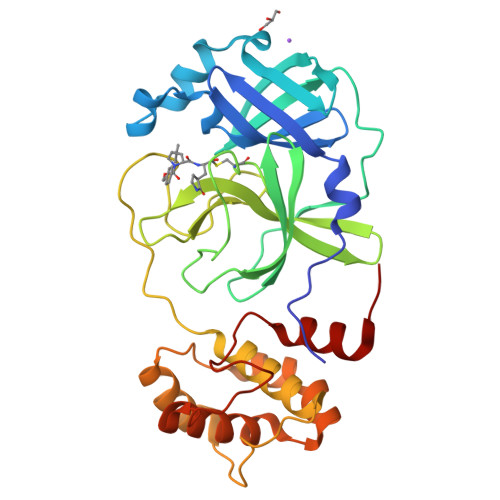
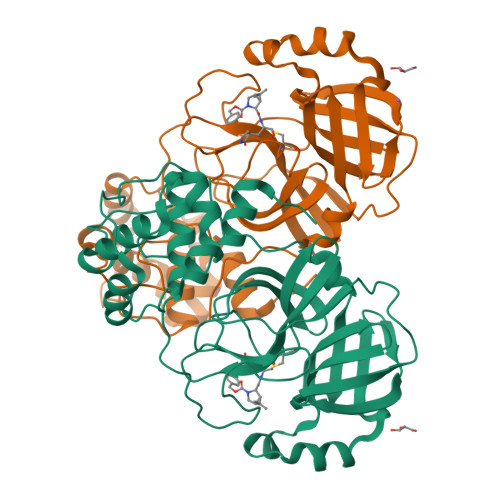
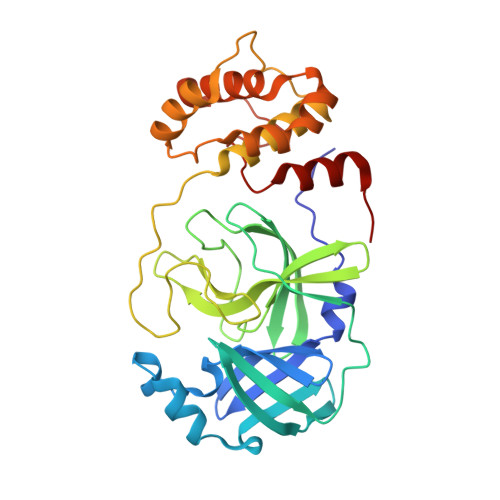
SARS-CoV-2 main protease (M pro ) is a cysteine protease that mediates the cleavage of viral polyproteins and is a validated antiviral drug target. M pro is highly conserved among all seven human coronaviruses, with certain M pro inhibitors having broad-spectrum antiviral activity. In this study, we designed two hybrid inhibitors UAWJ9-36-1 and UAWJ9-36-3 based on the superimposed X-ray crystal structures of SARS-CoV-2 M pro with GC-376, telaprevir, and boceprevir. Both UAWJ9-36-1 and UAWJ9-36-3 showed potent binding and enzymatic inhibition against the M pro 's from SARS-CoV-2, SARS-CoV, MERS-CoV, HCoV-OC43, HCoV-NL63, HCoV-229E, and HCoV-HKU1. Cell-based Flip-GFP M pro assay results show that UAWJ9-36-1 and UAWJ9-36-3 inhibited the intracellular protease activity of SARS-CoV-2 M pro . In addition, UAWJ9-36-1 and UAWJ9-36-3 had potent antiviral activity against SARS-CoV-2, HCoV-OC43, HCoV-NL63, and HCoV-229E, with UAWJ9-36-3 being more potent than GC-376 in inhibiting SARS-CoV-2. Selectivity profiling revealed that UAWJ9-36-1 and UAWJ9-36-3 had an improved selectivity index over that of GC-376 against host cysteine proteases calpain I and cathepsin L, but not cathepsin K. The X-ray crystal structures of SARS-CoV-2 M pro with UAWJ9-36-1 and UAWJ9-36-3 were both solved at 1.9 Å, which validated our design hypothesis. Overall, hybrid inhibitors UAWJ9-36-1 and UAWJ9-36-3 are promising candidates to be further developed as broad-spectrum coronavirus antivirals.
Organizational Affiliation:
Department of Pharmacology and Toxicology, College of Pharmacy, The University of Arizona, Tucson, Arizona 85721, United States.