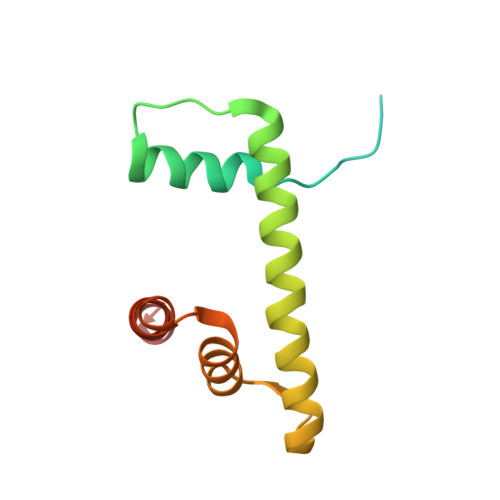
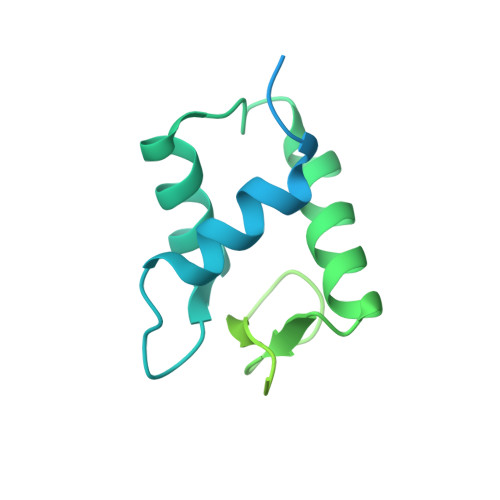
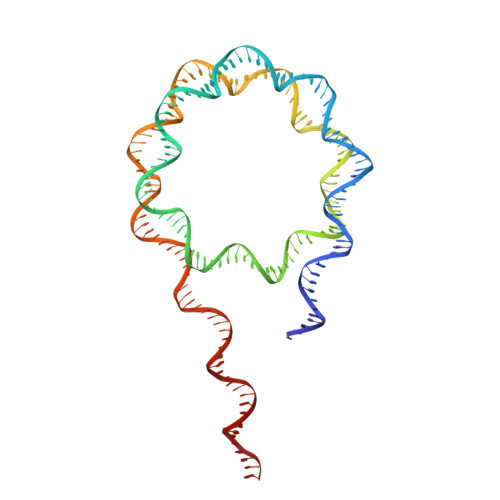
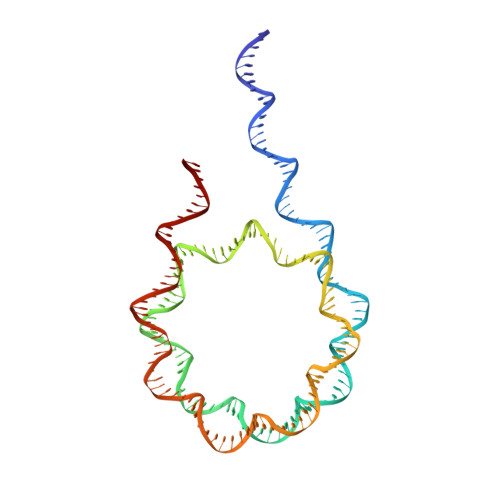
Histone H1 binding to nucleosome arrays depends on linker DNA length and trajectory.
Dombrowski, M., Engeholm, M., Dienemann, C., Dodonova, S., Cramer, P.(2022) Nat Struct Mol Biol 29: 493-501
- PubMed: 35581345
- DOI: https://doi.org/10.1038/s41594-022-00768-w
- Primary Citation of Related Structures:
7PET, 7PEU, 7PEV, 7PEW, 7PEX, 7PEY, 7PEZ, 7PF0, 7PF2, 7PF3, 7PF4, 7PF5, 7PF6, 7PFA, 7PFC, 7PFD, 7PFE, 7PFF, 7PFT, 7PFU, 7PFV, 7PFW, 7PFX - PubMed Abstract:
Throughout the genome, nucleosomes often form regular arrays that differ in nucleosome repeat length (NRL), occupancy of linker histone H1 and transcriptional activity. Here, we report cryo-EM structures of human H1-containing tetranucleosome arrays with four physiologically relevant NRLs. The structures show a zig-zag arrangement of nucleosomes, with nucleosomes 1 and 3 forming a stack. H1 binding to stacked nucleosomes depends on the NRL, whereas H1 always binds to the non-stacked nucleosomes 2 and 4. Short NRLs lead to altered trajectories of linker DNA, and these altered trajectories sterically impair H1 binding to the stacked nucleosomes in our structures. As the NRL increases, linker DNA trajectories relax, enabling H1 contacts and binding. Our results provide an explanation for why arrays with short NRLs are depleted of H1 and suited for transcription, whereas arrays with long NRLs show full H1 occupancy and can form transcriptionally silent heterochromatin regions.
Organizational Affiliation:
Department of Molecular Biology, Max Planck Institute for Multidisciplinary Sciences, Göttingen, Germany.