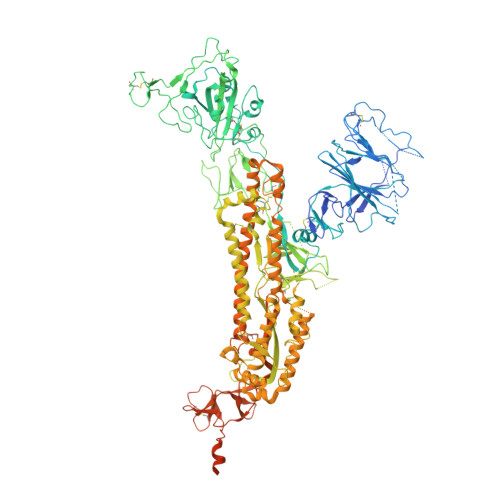
Evolution of the SARS-CoV-2 spike protein in the human host.
Wrobel, A.G., Benton, D.J., Roustan, C., Borg, A., Hussain, S., Martin, S.R., Rosenthal, P.B., Skehel, J.J., Gamblin, S.J.(2022) Nat Commun 13: 1178-1178
- PubMed: 35246509
- DOI: https://doi.org/10.1038/s41467-022-28768-w
- Primary Citation of Related Structures:
7R0Z, 7R10, 7R11, 7R12, 7R13, 7R14, 7R15, 7R16, 7R17, 7R18, 7R19, 7R1A, 7R1B - PubMed Abstract:
Recently emerged variants of SARS-CoV-2 contain in their surface spike glycoproteins multiple substitutions associated with increased transmission and resistance to neutralising antibodies. We have examined the structure and receptor binding properties of spike proteins from the B.1.1.7 (Alpha) and B.1.351 (Beta) variants to better understand the evolution of the virus in humans. Spikes of both variants have the same mutation, N501Y, in the receptor-binding domains. This substitution confers tighter ACE2 binding, dependent on the common earlier substitution, D614G. Each variant spike has acquired other key changes in structure that likely impact virus pathogenesis. The spike from the Alpha variant is more stable against disruption upon binding ACE2 receptor than all other spikes studied. This feature is linked to the acquisition of a more basic substitution at the S1-S2 furin site (also observed for the variants of concern Delta, Kappa, and Omicron) which allows for near-complete cleavage. In the Beta variant spike, the presence of a new substitution, K417N (also observed in the Omicron variant), in combination with the D614G, stabilises a more open spike trimer, a conformation required for receptor binding. Our observations suggest ways these viruses have evolved to achieve greater transmissibility in humans.
Organizational Affiliation:
Structural Biology of Disease Processes Laboratory, NW1 1AT, London, UK. antoni.wrobel@crick.ac.uk.