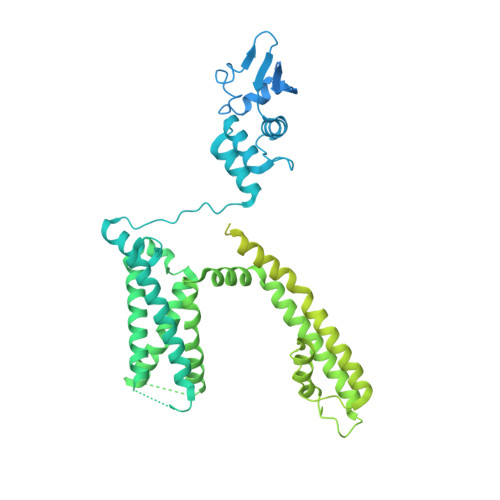
Structures of the T cell potassium channel Kv1.3 with immunoglobulin modulators.
Selvakumar, P., Fernandez-Marino, A.I., Khanra, N., He, C., Paquette, A.J., Wang, B., Huang, R., Smider, V.V., Rice, W.J., Swartz, K.J., Meyerson, J.R.(2022) Nat Commun 13: 3854-3854
- PubMed: 35788586
- DOI: https://doi.org/10.1038/s41467-022-31285-5
- Primary Citation of Related Structures:
7SSV, 7SSX, 7SSY, 7SSZ, 8DFL - PubMed Abstract:
The Kv1.3 potassium channel is expressed abundantly on activated T cells and mediates the cellular immune response. This role has made the channel a target for therapeutic immunomodulation to block its activity and suppress T cell activation. Here, we report structures of human Kv1.3 alone, with a nanobody inhibitor, and with an antibody-toxin fusion blocker. Rather than block the channel directly, four copies of the nanobody bind the tetramer's voltage sensing domains and the pore domain to induce an inactive pore conformation. In contrast, the antibody-toxin fusion docks its toxin domain at the extracellular mouth of the channel to insert a critical lysine into the pore. The lysine stabilizes an active conformation of the pore yet blocks ion permeation. This study visualizes Kv1.3 pore dynamics, defines two distinct mechanisms to suppress Kv1.3 channel activity with exogenous inhibitors, and provides a framework to aid development of emerging T cell immunotherapies.
Organizational Affiliation:
Department of Physiology and Biophysics, Weill Cornell Medical College, New York, NY, USA.