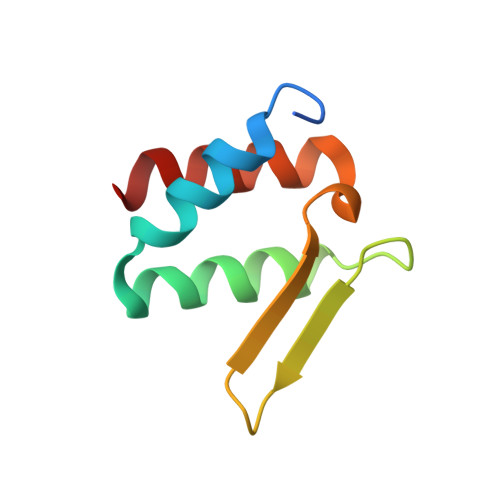
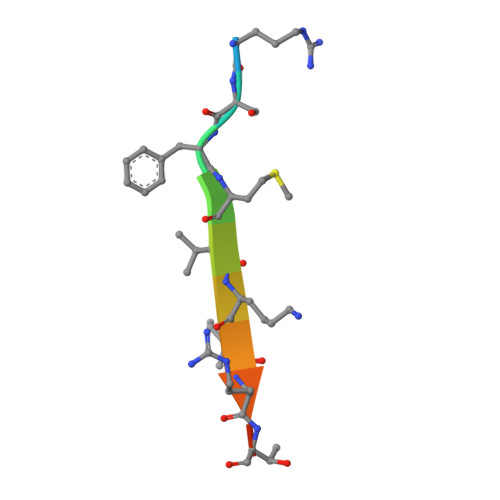
Taf2 mediates DNA binding of Taf14.
Klein, B.J., Feigerle, J.T., Zhang, J., Ebmeier, C.C., Fan, L., Singh, R.K., Wang, W.W., Schmitt, L.R., Lee, T., Hansen, K.C., Liu, W.R., Wang, Y.X., Strahl, B.D., Anthony Weil, P., Kutateladze, T.G.(2022) Nat Commun 13: 3177-3177
- PubMed: 35676274
- DOI: https://doi.org/10.1038/s41467-022-30937-w
- Primary Citation of Related Structures:
7UHE - PubMed Abstract:
The assembly and function of the yeast general transcription factor TFIID complex requires specific contacts between its Taf14 and Taf2 subunits, however, the mechanism underlying these contacts remains unclear. Here, we determined the molecular and structural basis by which the YEATS and ET domains of Taf14 bind to the C-terminal tail of Taf2 and identified a unique DNA-binding activity of the linker region connecting the two domains. We show that in the absence of ligands the linker region of Taf14 is occluded by the surrounding domains, and therefore the DNA binding function of Taf14 is autoinhibited. Binding of Taf2 promotes a conformational rearrangement in Taf14, resulting in a release of the linker for the engagement with DNA and the nucleosome. Genetic in vivo data indicate that the association of Taf14 with both Taf2 and DNA is essential for transcriptional regulation. Our findings provide a basis for deciphering the role of individual TFIID subunits in mediating gene transcription.
Organizational Affiliation:
Department of Pharmacology, University of Colorado School of Medicine, Aurora, CO, 80045, USA.