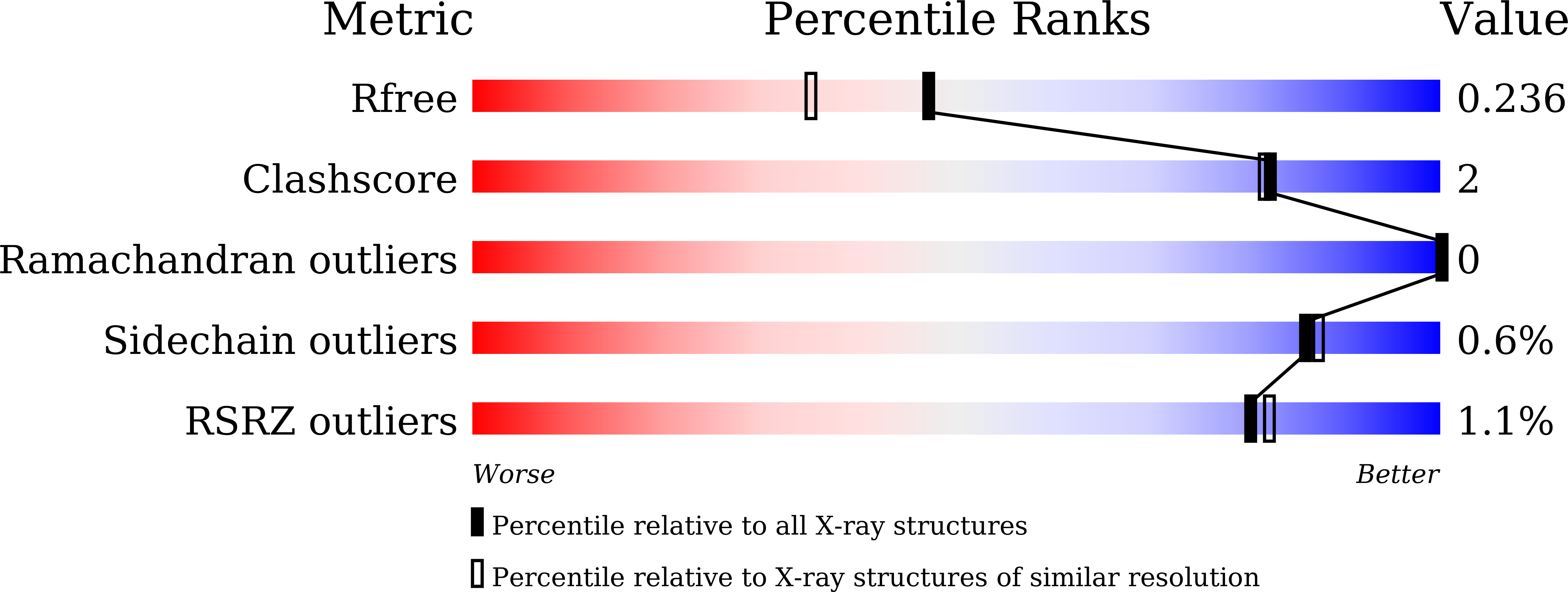
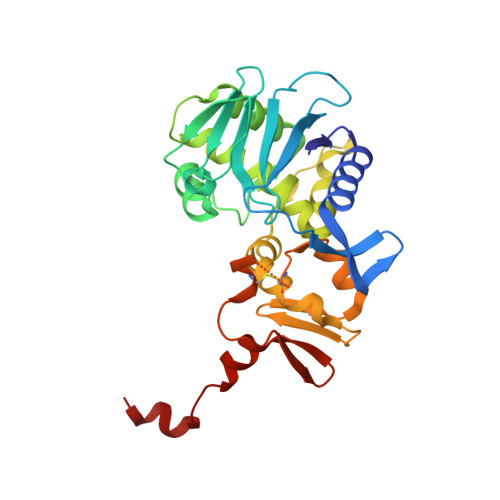
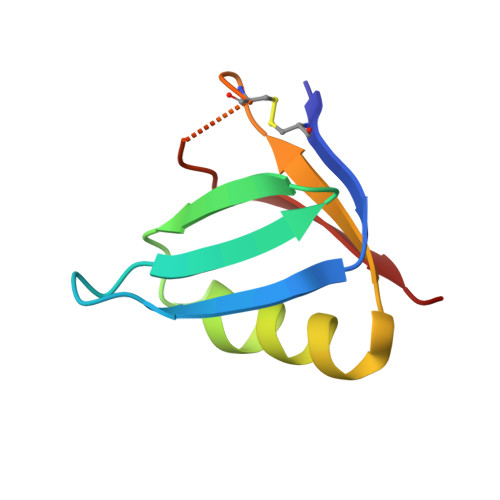
A unique peptide-based pharmacophore identifies an inhibitory compound against the A-subunit of Shiga toxin.
Watanabe-Takahashi, M., Senda, M., Yoshino, R., Hibino, M., Hama, S., Terada, T., Shimizu, K., Senda, T., Nishikawa, K.(2022) Sci Rep 12: 11443-11443
- PubMed: 35794188
- DOI: https://doi.org/10.1038/s41598-022-15316-1
- Primary Citation of Related Structures:
7VHC, 7VHD, 7VHE, 7VHF - PubMed Abstract:
Shiga toxin (Stx), a major virulence factor of enterohemorrhagic Escherichia coli (EHEC), can cause fatal systemic complications. Recently, we identified a potent inhibitory peptide that binds to the catalytic A-subunit of Stx. Here, using biochemical structural analysis and X-ray crystallography, we determined a minimal essential peptide motif that occupies the catalytic cavity and is required for binding to the A-subunit of Stx2a, a highly virulent Stx subtype. Molecular dynamics simulations also identified the same motif and allowed determination of a unique pharmacophore for A-subunit binding. Notably, a series of synthetic peptides containing the motif efficiently inhibit Stx2a. In addition, pharmacophore screening and subsequent docking simulations ultimately identified nine Stx2a-interacting molecules out of a chemical compound database consisting of over 7,400,000 molecules. Critically, one of these molecules markedly inhibits Stx2a both in vitro and in vivo, clearly demonstrating the significance of the pharmacophore for identifying therapeutic agents against EHEC infection.
Organizational Affiliation:
Department of Molecular Life Sciences, Graduate School of Life and Medical Sciences, Doshisha University, Kyoto, Japan.