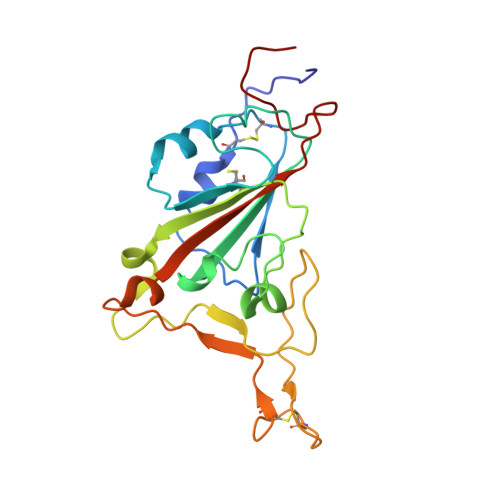
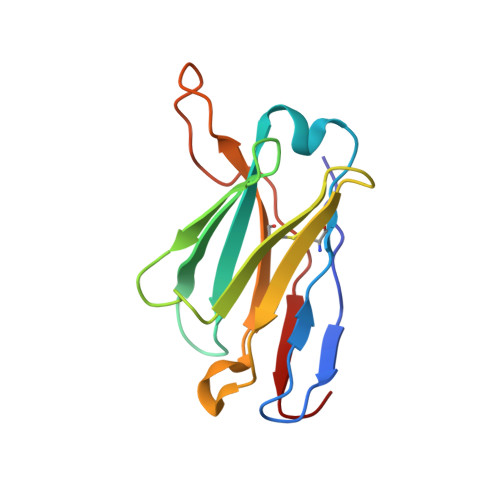
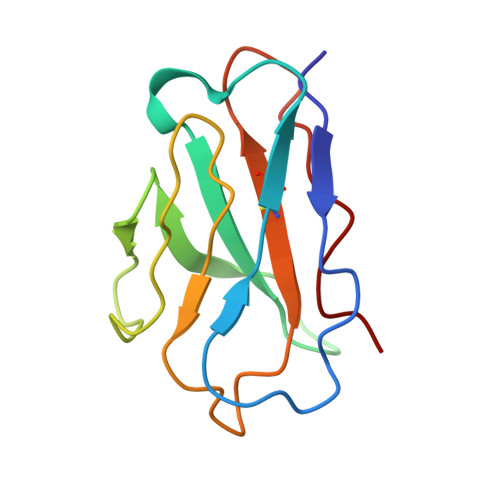
Immunoglobulin germline gene polymorphisms influence the function of SARS-CoV-2 neutralizing antibodies.
Pushparaj, P., Nicoletto, A., Sheward, D.J., Das, H., Castro Dopico, X., Perez Vidakovics, L., Hanke, L., Chernyshev, M., Narang, S., Kim, S., Fischbach, J., Ekstrom, S., McInerney, G., Hallberg, B.M., Murrell, B., Corcoran, M., Karlsson Hedestam, G.B.(2023) Immunity 56: 193-206.e7
- PubMed: 36574772
- DOI: https://doi.org/10.1016/j.immuni.2022.12.005
- Primary Citation of Related Structures:
8A94, 8A95, 8A96, 8A99 - PubMed Abstract:
The human immunoglobulin heavy-chain (IGH) locus is exceptionally polymorphic, with high levels of allelic and structural variation. Thus, germline IGH genotypes are personal, which may influence responses to infection and vaccination. For an improved understanding of inter-individual differences in antibody responses, we isolated SARS-CoV-2 spike-specific monoclonal antibodies from convalescent health care workers, focusing on the IGHV1-69 gene, which has the highest level of allelic variation of all IGHV genes. The IGHV1-69 ∗ 20-using CAB-I47 antibody and two similar antibodies isolated from an independent donor were critically dependent on allele usage. Neutralization was retained when reverting the V region to the germline IGHV1-69 ∗ 20 allele but lost when reverting to other IGHV1-69 alleles. Structural data confirmed that two germline-encoded polymorphisms, R50 and F55, in the IGHV1-69 gene were required for high-affinity receptor-binding domain interaction. These results demonstrate that polymorphisms in IGH genes can influence the function of SARS-CoV-2 neutralizing antibodies.
Organizational Affiliation:
Department of Microbiology, Tumor and Cell Biology, Karolinska Institutet, 171 77 Stockholm, Sweden.