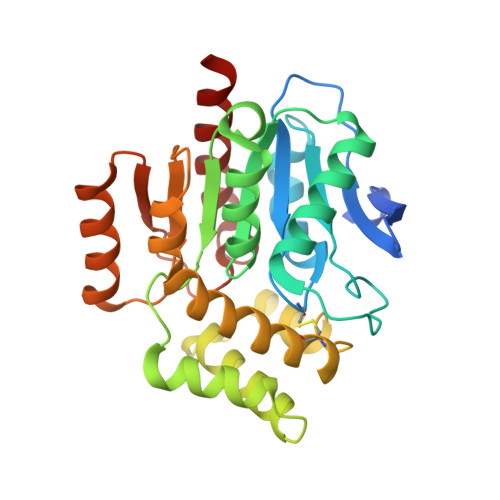
Evolutionary adaptation from hydrolytic to oxygenolytic catalysis at the alpha / beta-hydrolase fold.
Bui, S., Gil-Guerrero, S., van der Linden, P., Carpentier, P., Ceccarelli, M., Jambrina, P.G., Steiner, R.A.(2023) Chem Sci 14: 10547-10560
- PubMed: 37799987
- DOI: https://doi.org/10.1039/d3sc03044j
- Primary Citation of Related Structures:
7OJM, 7OKZ, 8A97, 8ORO, 8OXN, 8OXT - PubMed Abstract:
Protein fold adaptation to novel enzymatic reactions is a fundamental evolutionary process. Cofactor-independent oxygenases degrading N -heteroaromatic substrates belong to the α/β-hydrolase (ABH) fold superfamily that typically does not catalyze oxygenation reactions. Here, we have integrated crystallographic analyses under normoxic and hyperoxic conditions with molecular dynamics and quantum mechanical calculations to investigate its prototypic 1- H -3-hydroxy-4-oxoquinaldine 2,4-dioxygenase (HOD) member. O 2 localization to the "oxyanion hole", where catalysis occurs, is an unfavorable event and the direct competition between dioxygen and water for this site is modulated by the "nucleophilic elbow" residue. A hydrophobic pocket that overlaps with the organic substrate binding site can act as a proximal dioxygen reservoir. Freeze-trap pressurization allowed the structure of the ternary complex with a substrate analogue and O 2 bound at the oxyanion hole to be determined. Theoretical calculations reveal that O 2 orientation is coupled to the charge of the bound organic ligand. When 1- H -3-hydroxy-4-oxoquinaldine is uncharged, O 2 binds with its molecular axis along the ligand's C2-C4 direction in full agreement with the crystal structure. Substrate activation triggered by deprotonation of its 3-OH group by the His-Asp dyad, rotates O 2 by approximately 60°. This geometry maximizes the charge transfer between the substrate and O 2 , thus weakening the double bond of the latter. Electron density transfer to the O 2 (π*) orbital promotes the formation of the peroxide intermediate via intersystem crossing that is rate-determining. Our work provides a detailed picture of how evolution has repurposed the ABH-fold architecture and its simple catalytic machinery to accomplish metal-independent oxygenation.
Organizational Affiliation:
Randall Centre for Cell and Molecular Biophysics, King's College London London SE1 1UL UK roberto.steiner@kcl.ac.uk.