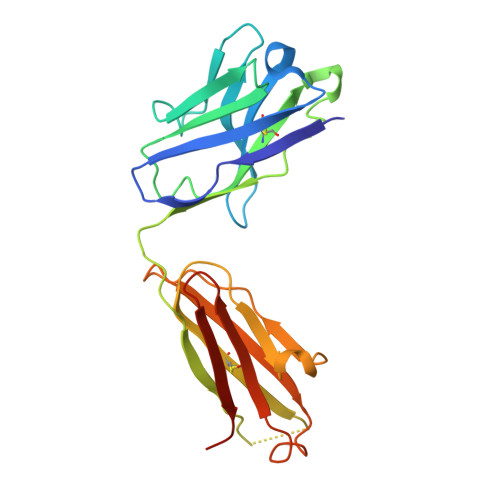
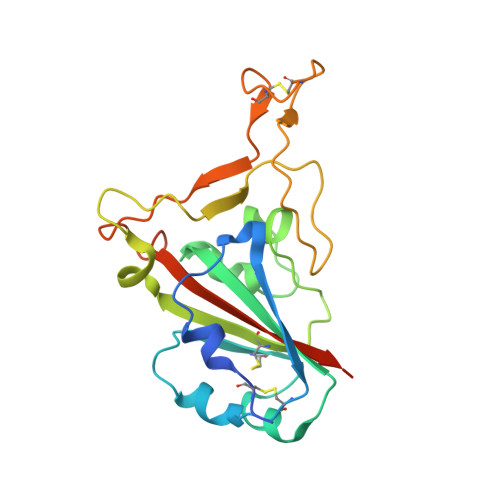
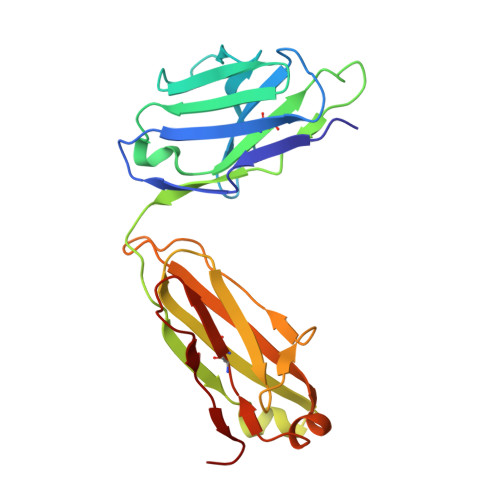
Emerging variants develop total escape from potent monoclonal antibodies induced by BA.4/5 infection.
Liu, C., Das, R., Dijokaite-Guraliuc, A., Zhou, D., Mentzer, A.J., Supasa, P., Selvaraj, M., Duyvesteyn, H.M.E., Ritter, T.G., Temperton, N., Klenerman, P., Dunachie, S.J., Paterson, N.G., Williams, M.A., Hall, D.R., Fry, E.E., Mongkolsapaya, J., Ren, J., Stuart, D.I., Screaton, G.R.(2024) Nat Commun 15: 3284-3284
- PubMed: 38627386
- DOI: https://doi.org/10.1038/s41467-024-47393-3
- Primary Citation of Related Structures:
8CBD, 8CBE, 8CBF, 8CMA, 8QZR - PubMed Abstract:
The rapid evolution of SARS-CoV-2 is driven in part by a need to evade the antibody response in the face of high levels of immunity. Here, we isolate spike (S) binding monoclonal antibodies (mAbs) from vaccinees who suffered vaccine break-through infections with Omicron sub lineages BA.4 or BA.5. Twenty eight potent antibodies are isolated and characterised functionally, and in some cases structurally. Since the emergence of BA.4/5, SARS-CoV-2 has continued to accrue mutations in the S protein, to understand this we characterize neutralization of a large panel of variants and demonstrate a steady attrition of neutralization by the panel of BA.4/5 mAbs culminating in total loss of function with recent XBB.1.5.70 variants containing the so-called 'FLip' mutations at positions 455 and 456. Interestingly, activity of some mAbs is regained on the recently reported variant BA.2.86.
Organizational Affiliation:
Chinese Academy of Medical Science (CAMS) Oxford Institute (COI), University of Oxford, Oxford, UK.