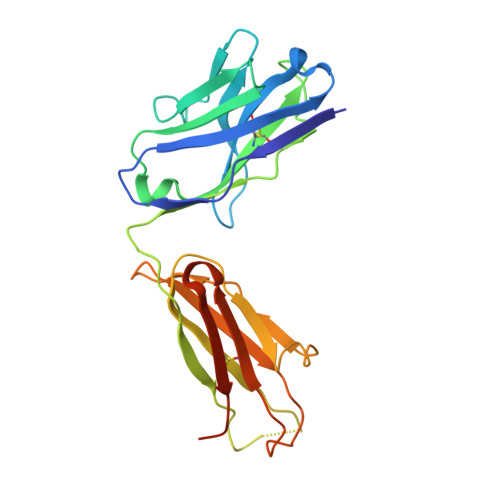
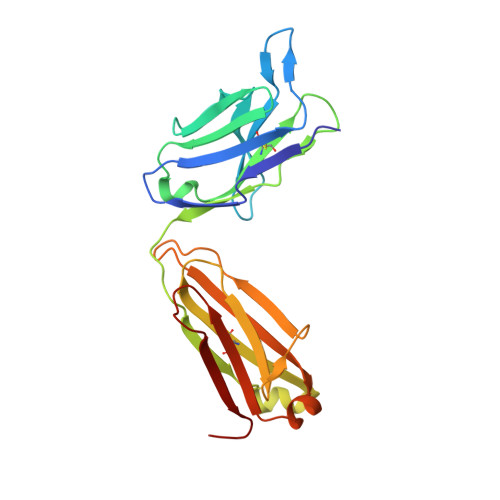
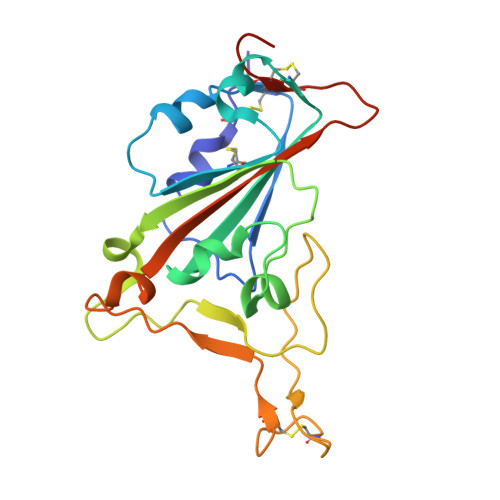
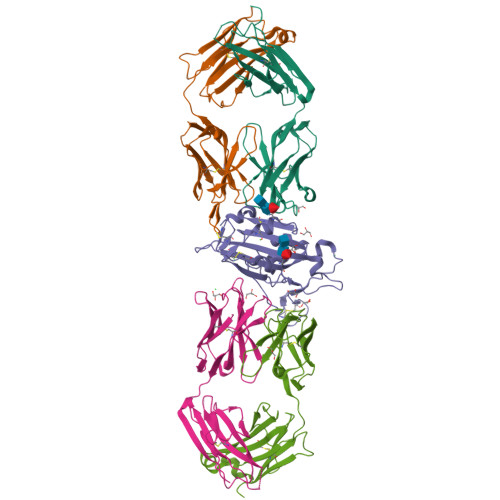Affinity maturation endows potent activity onto class 6 SARS-CoV-2 broadly neutralizing antibodies.
Mazigi, O., Langley, D.B., Henry, J.Y., Burnett, D.L., Sobti, M., Walker, G.J., Rouet, R., Balachandran, H., Lenthall, H., Jackson, J., Ubiparipovic, S., Schofield, P., Brown, S.H.J., Schulz, S.R., Hoffmann, M., Pohlmann, S., Post, J., Martinello, M., Ahlenstiel, G., Kelleher, A., Rawlinson, W.D., Turville, S.G., Bull, R.A., Stewart, A.G., Jack, H.M., Goodnow, C.C., Christ, D.(2025) Proc Natl Acad Sci U S A 122: e2417544121-e2417544121
- PubMed: 39746041
- DOI: https://doi.org/10.1073/pnas.2417544121
- Primary Citation of Related Structures:
8CWI, 8CWJ, 8CWK - PubMed Abstract:
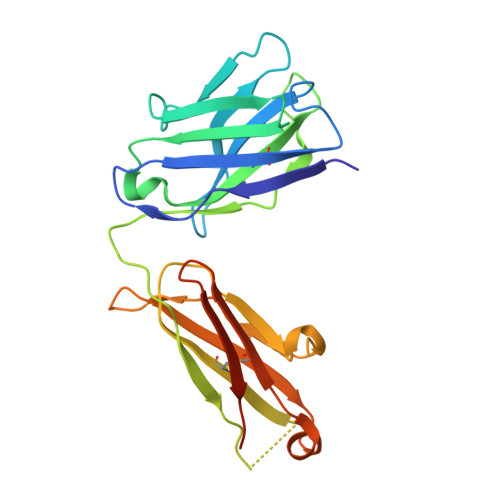
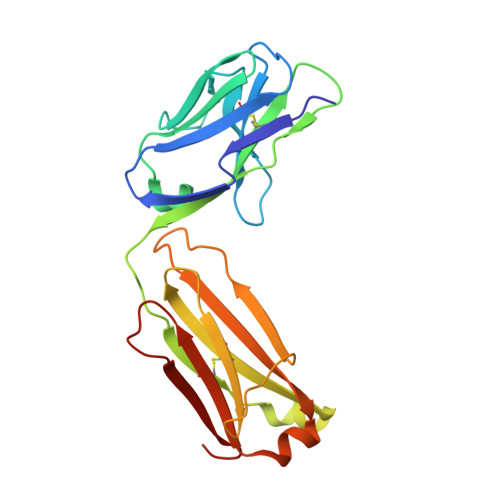
The emergence of SARS-CoV-2 variants of concern (VOCs) has greatly diminished the neutralizing activity of previously FDA-approved monoclonal antibodies (mAbs), including that of antibody cocktails and of first-generation broadly neutralizing antibodies such as S309 (Sotrovimab). In contrast, antibodies targeting cryptic conformational epitopes of the receptor binding domain (RBD) have demonstrated broad activity against emerging variants, but exert only moderate neutralizing activity, which has so far hindered clinical development. Here, we utilize in vitro display technology to identify and affinity-mature antibodies targeting the cryptic class 6 epitope, accessible only in the "up" conformation of the SARS-CoV-2 spike trimer. Increasing antibody affinity into the low picomolar range endowed potent neutralization of VOCs and protection of hACE2 mice from viral challenge. Cryoelectron microscopy and crystal structures of two affinity-matured antibodies (4C12-B12 and 4G1-C2) in complex with RBD highlighted binding modes and epitopes distal from mutational hotspots commonly overserved in VOCs, providing direct structural insights into the observed mutational resistance. Moreover, we further demonstrate that antibodies targeting the class 6 epitope, rather than being an artifact of in vitro selection, are common in the IgG1 + memory B cell repertoire of convalescent patients and can be induced in human antibody V-gene transgenic mice through immunization. Our results highlight the importance of very high (picomolar) affinity in the development of neutralizing antibodies and vaccines and suggest an affinity threshold in the provision of broad and long-lasting immunity against SARS-CoV-2.
Organizational Affiliation:
Garvan Institute of Medical Research, Sydney, NSW 2010, Australia.