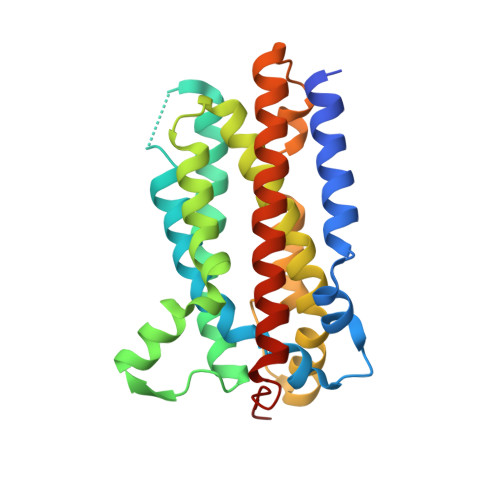
The structural and functional investigation into an unusual nitrile synthase.
Li, H., Huang, J.W., Dai, L., Zheng, H., Dai, S., Zhang, Q., Yao, L., Yang, Y., Yang, Y., Min, J., Guo, R.T., Chen, C.C.(2023) Nat Commun 14: 7425-7425
- PubMed: 37973794
- DOI: https://doi.org/10.1038/s41467-023-43285-0
- Primary Citation of Related Structures:
8JI2, 8JI3, 8JI4, 8JI5, 8JI6, 8JI7 - PubMed Abstract:
The biosynthesis of neurotoxin aetokthonotoxin (AETX) that features a unique structure of pentabrominated biindole nitrile involves a first-of-its-kind nitrile synthase termed AetD, an enzyme that shares very low sequence identity to known structures and catalyzes an unprecedented mechanism. In this study, we resolve the crystal structure of AetD in complex with the substrate 5,7-di-Br-L-Trp. AetD adopts the heme oxygenase like fold and forms a hydrophobic cavity within a helical bundle to accommodate the indole moiety. A diiron cluster comprising two irons that serves as a catalytic center binds to the carboxyl O and the amino N of the substrate. Notably, we demonstrate that the AetD-catalyzed reaction is independent of the bromination of the substrate and also solved crystal structures of AetD in complex with 5-Br-L-Trp and L-Trp. Altogether, the present study reveals the substrate-binding pattern and validates the diiron cluster-comprising active center of AetD, which should provide important basis to support the mechanistic investigations into this class of nitrile synthase.
Organizational Affiliation:
State Key Laboratory of Biocatalysis and Enzyme Engineering, Hubei Hongshan Laboratory, Hubei Collaborative Innovation Center for Green Transformation of Bio-Resources, Hubei Key Laboratory of Industrial Biotechnology, School of Life Sciences, Hubei University, Wuhan, 430062, PR China.