Structure of active state MC4R in complex with a potent ligand mimicking nanobody
Busch, A., Jaakola, V.-P., Masiulis, S.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Guanine nucleotide-binding protein G(s) subunit alpha isoforms short | A [auth B] | 394 | Homo sapiens | Mutation(s): 9 Gene Names: GNAS EC: 3.6.5 Membrane Entity: Yes | 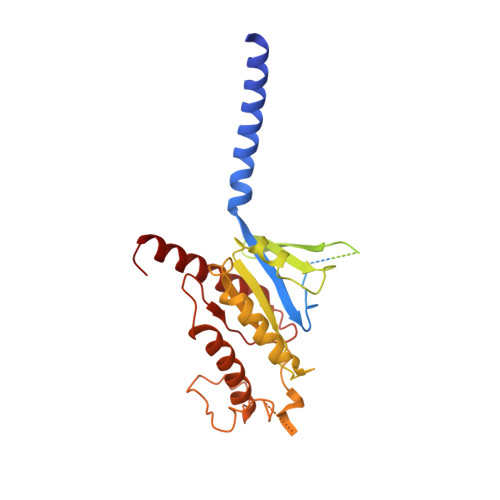 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P63092 (Homo sapiens) Explore P63092 Go to UniProtKB: P63092 | |||||
PHAROS: P63092 GTEx: ENSG00000087460 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P63092 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Guanine nucleotide-binding protein G(I)/G(S)/G(T) subunit beta-1 | B [auth C] | 350 | Homo sapiens | Mutation(s): 0 Gene Names: GNB1 |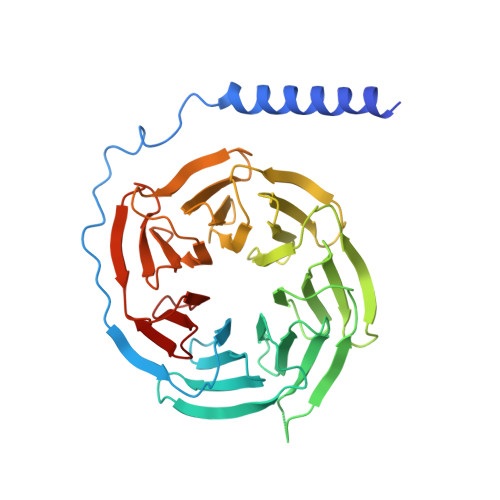  |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P62873 (Homo sapiens) Explore P62873 Go to UniProtKB: P62873 | |||||
PHAROS: P62873 GTEx: ENSG00000078369 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P62873 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 | C [auth G] | 71 | Homo sapiens | Mutation(s): 0 Gene Names: GNG2 Membrane Entity: Yes | 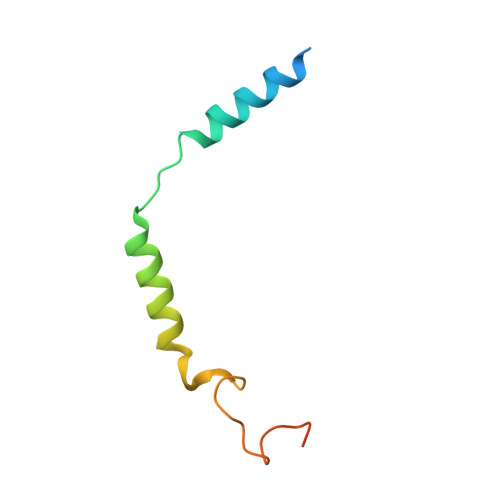 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P59768 (Homo sapiens) Explore P59768 Go to UniProtKB: P59768 | |||||
PHAROS: P59768 GTEx: ENSG00000186469 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P59768 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Nanobody-35 | D [auth N] | 138 | Lama glama | Mutation(s): 0 | 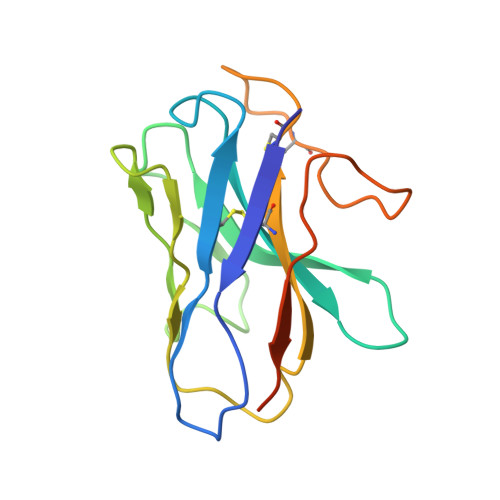 |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Soluble cytochrome b562,Melanocortin receptor 4,Red fluorescent protein drFP583 | E [auth A] | 729 | Escherichia coli, Homo sapiens, Discosoma sp. This entity is chimeric | Mutation(s): 32 Gene Names: cybC, MC4R Membrane Entity: Yes | 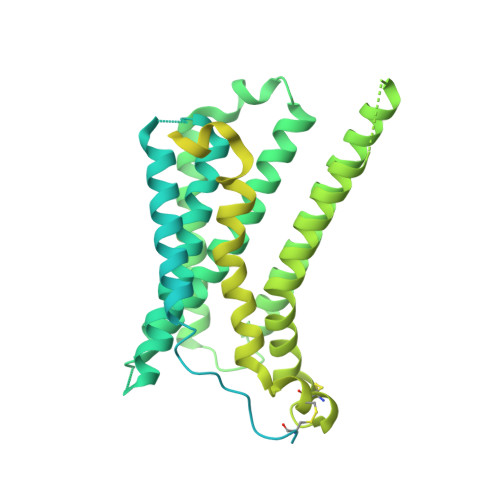 |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P0ABE7 (Escherichia coli) Explore P0ABE7 Go to UniProtKB: P0ABE7 | |||||
Find proteins for Q9U6Y8 (Discosoma sp.) Explore Q9U6Y8 Go to UniProtKB: Q9U6Y8 | |||||
Find proteins for P32245 (Homo sapiens) Explore P32245 Go to UniProtKB: P32245 | |||||
PHAROS: P32245 GTEx: ENSG00000166603 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Groups | Q9U6Y8P0ABE7P32245 | ||||
Sequence AnnotationsExpand | |||||
| |||||
Entity ID: 6 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| pN162 | F [auth D] | 117 | Lama glama | Mutation(s): 0 |  |
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Funding Organization | Location | Grant Number |
|---|---|---|
| Other government | Belgium | 2016-R-49 |
| Other government | Belgium | HBC_2020.2042 |