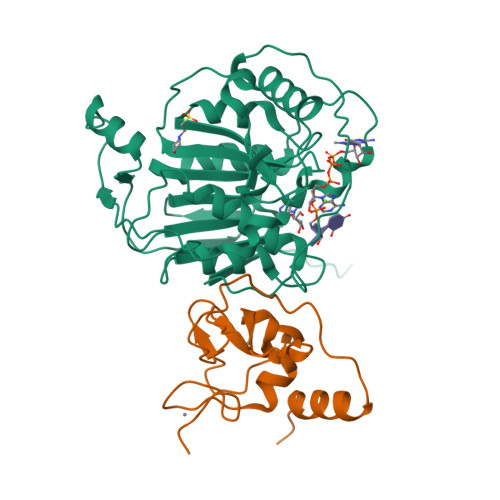
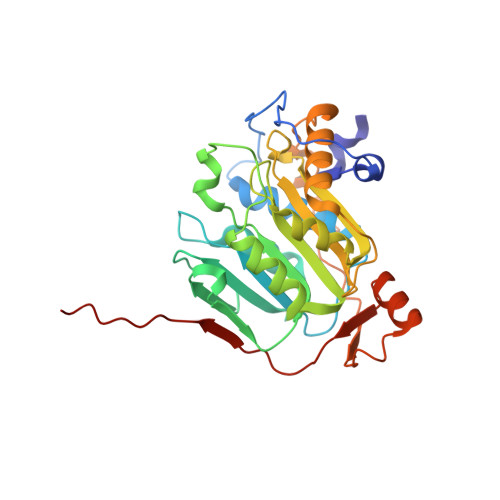
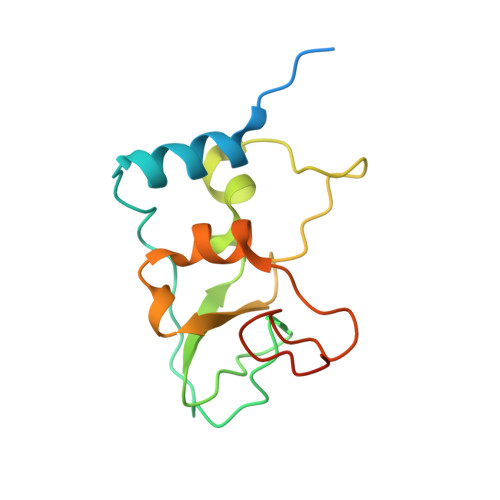
SARS-CoV-2 methyltransferase nsp10-16 in complex with natural and drug-like purine analogs for guiding structure-based drug discovery
Kremling, V., Falke, S., Fernandez-Garcia, Y., Ehrt, C., Kiene, A., Klopprogge, B., Scheer, E., Barthels, F., Middendorf, P., Kuhn, S., Gunther, S., Rarey, M., Chapman, H.N., Oberthur, D., Sprenger, J.(2024) Elife