Structure of Trek-1(S131C mutant) with ML335
Mondal, A., Lee, H., Minor, D.L.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Potassium channel subfamily K member 2 | 287 | Mus musculus | Mutation(s): 14 Gene Names: Kcnk2 Membrane Entity: Yes | 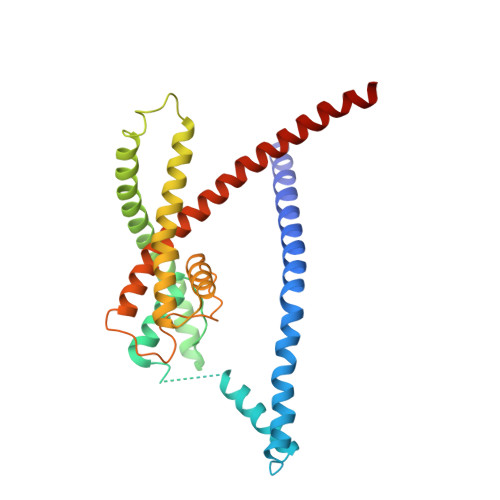 | |
UniProt & NIH Common Fund Data Resources | |||||
Find proteins for P97438 (Mus musculus) Explore P97438 Go to UniProtKB: P97438 | |||||
IMPC: MGI:109366 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P97438 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 6 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 16C Query on 16C | C [auth A] | N-((E,2S,3R)-1,3-DIHYDROXYOCTADEC-4-EN-2-YL)PALMITAMIDE C34 H67 N O3 YDNKGFDKKRUKPY-TURZORIXSA-N |  | ||
| WUZ (Subject of Investigation/LOI) Query on WUZ | G [auth A], U [auth B] | N-[(2,4-dichlorophenyl)methyl]-4-[(3R)-3-methyl-2,5-dioxopyrrolidin-1-yl]benzamide C19 H16 Cl2 N2 O3 XQJYUMZZHNFDCF-LLVKDONJSA-N |  | ||
| R16 Query on R16 | L [auth A], X [auth B], Y [auth B] | HEXADECANE C16 H34 DCAYPVUWAIABOU-UHFFFAOYSA-N |  | ||
| D10 Query on D10 | AA [auth B] D [auth A] E [auth A] F [auth A] H [auth A] | DECANE C10 H22 DIOQZVSQGTUSAI-UHFFFAOYSA-N |  | ||
| CD Query on CD | M [auth A], N [auth A], Z [auth B] | CADMIUM ION Cd WLZRMCYVCSSEQC-UHFFFAOYSA-N |  | ||
| K Query on K | I [auth A], J [auth A], K [auth A], V [auth B], W [auth B] | POTASSIUM ION K NPYPAHLBTDXSSS-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 67.213 | α = 90 |
| b = 120.568 | β = 90 |
| c = 128.613 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| XDS | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Institutes of Health/National Institute of Mental Health (NIH/NIMH) | United States | R01-MH116278 |
| National Institutes of Health/National Institute of Mental Health (NIH/NIMH) | United States | R01-MH093603 |