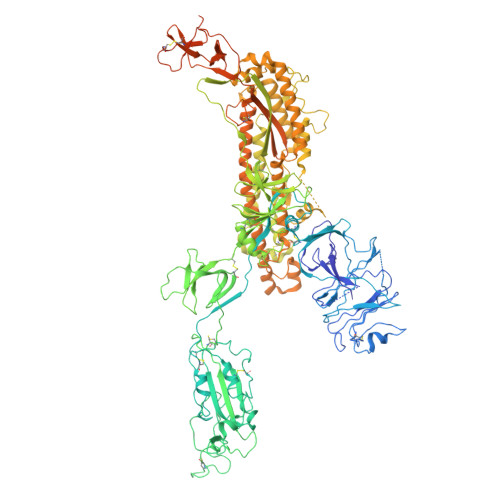
Structural basis for receptor-binding domain mobility of the spike in SARS-CoV-2 BA.2.86 and JN.1.
Yajima, H., Anraku, Y., Kaku, Y., Kimura, K.T., Plianchaisuk, A., Okumura, K., Nakada-Nakura, Y., Atarashi, Y., Hemmi, T., Kuroda, D., Takahashi, Y., Kita, S., Sasaki, J., Sumita, H., Ito, J., Maenaka, K., Sato, K., Hashiguchi, T.(2024) Nat Commun 15: 8574-8574
- PubMed: 39375326
- DOI: https://doi.org/10.1038/s41467-024-52808-2
- Primary Citation of Related Structures:
8WXL, 8XUX, 8XUY, 8XUZ, 8XV0, 8XV1, 8XVM, 9IU1 - PubMed Abstract:
Since 2019, SARS-CoV-2 has undergone mutations, resulting in pandemic and epidemic waves. The SARS-CoV-2 spike protein, crucial for cellular entry, binds to the ACE2 receptor exclusively when its receptor-binding domain (RBD) adopts the up-conformation. However, whether ACE2 also interacts with the RBD in the down-conformation to facilitate the conformational shift to RBD-up remains unclear. Herein, we present the structures of the BA.2.86 and the JN.1 spike proteins bound to ACE2. Notably, we successfully observed the ACE2-bound down-RBD, indicating an intermediate structure before the RBD-up conformation. The wider and mobile angle of RBDs in the up-state provides space for ACE2 to interact with the down-RBD, facilitating the transition to the RBD-up state. The K356T, but not N354-linked glycan, contributes to both of infectivity and neutralizing-antibody evasion in BA.2.86. These structural insights the spike-protein dynamics would help understand the mechanisms underlying SARS-CoV-2 infection and its neutralization.
Organizational Affiliation:
Laboratory of Medical Virology, Institute for Life and Medical Sciences, Kyoto University, Kyoto, Japan.