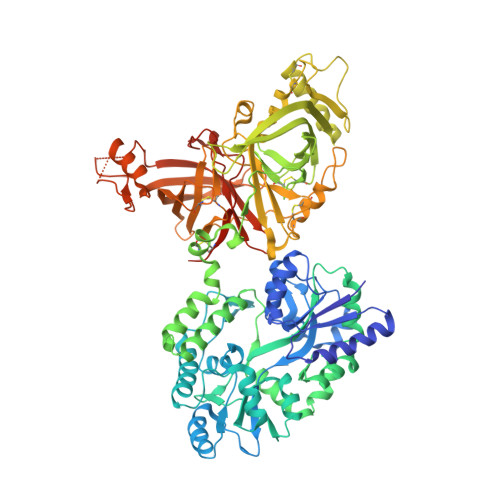
Structure of WNT inhibitor adenomatosis polyposis coli down-regulated 1 (APCDD1), a cell-surface lipid-binding protein.
Hsieh, F.L., Chang, T.H., Gabelli, S.B., Nathans, J.(2023) Proc Natl Acad Sci U S A 120: e2217096120-e2217096120
- PubMed: 37155902
- DOI: https://doi.org/10.1073/pnas.2217096120
- Primary Citation of Related Structures:
8E0P, 8E0R, 8E0W - PubMed Abstract:
Diverse extracellular proteins negatively regulate WNT signaling. One such regulator is adenomatosis polyposis coli down-regulated 1 (APCDD1), a conserved single-span transmembrane protein. In response to WNT signaling in a variety of tissues, APCDD1 transcripts are highly up-regulated. We have determined the three-dimensional structure of the extracellular domain of APCDD1, and this structure reveals an unusual architecture consisting of two closely apposed β-barrel domains (ABD1 and ABD2). ABD2, but not ABD1, has a large hydrophobic pocket that accommodates a bound lipid. The APCDD1 ECD can also bind to WNT7A, presumably via its covalently bound palmitoleate, a modification that is common to all WNTs and is essential for signaling. This work suggests that APCDD1 functions as a negative feedback regulator by titrating WNT ligands at the surface of responding cells.
Organizational Affiliation:
Department of Molecular Biology and Genetics, Johns Hopkins University School of Medicine, Baltimore, MD 21205.