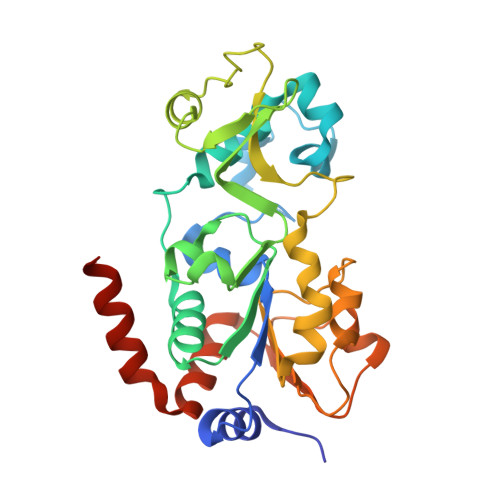
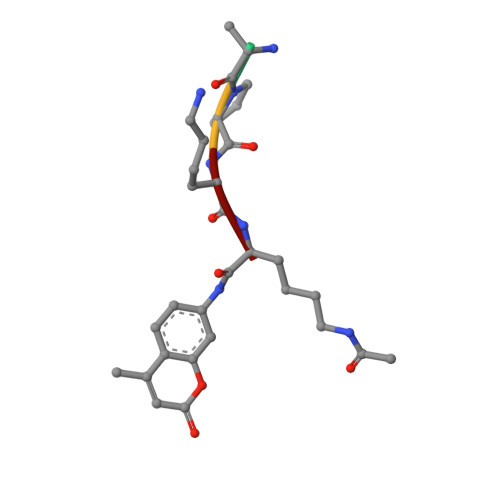
Computationally Driven Discovery and Characterization of SIRT3 Activating Compounds that Fully Recover Catalytic Activity under NAD+ Depletion
Chakrabarti, R., Ghosh, A., Guan, X., Upadhyay, A., Dumpati, R.K., Munshi, S., Roy, S., Chall, S., Rahnamoun, A., Reverdy, C., Errasti, G., Delacroix, T.(2023) bioRxiv