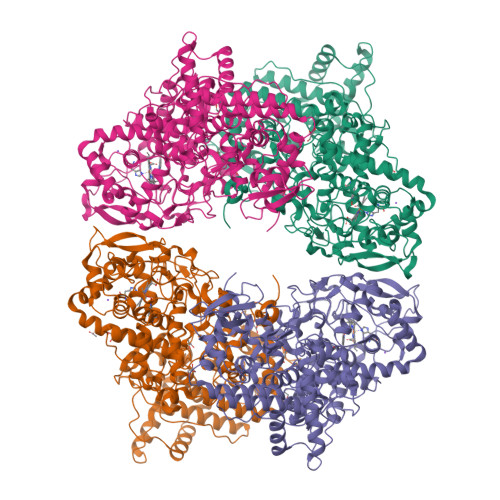
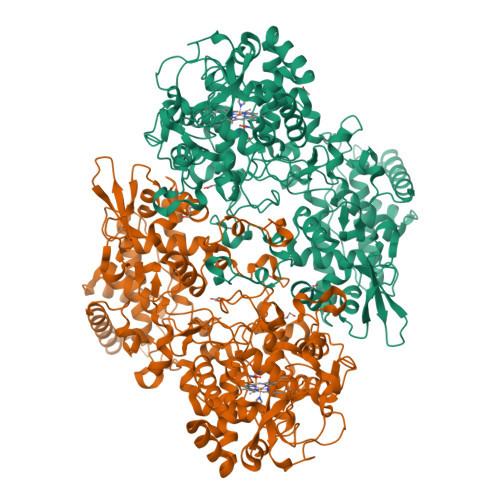
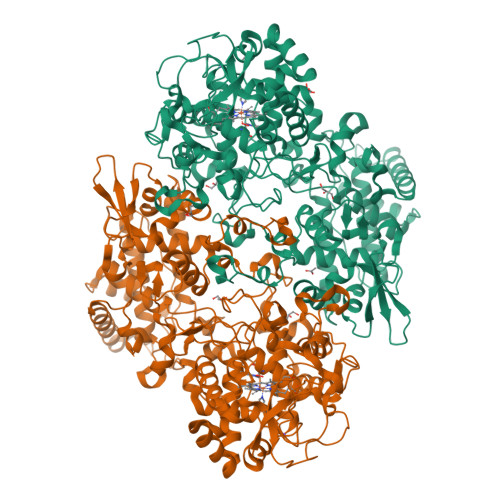
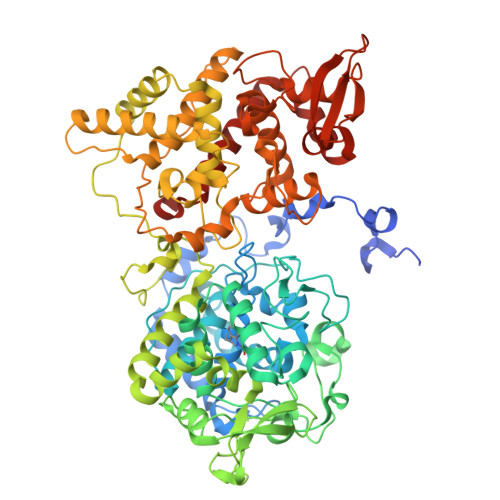
Indole N-Linked Hydroperoxyl Adduct of Protein-Derived Cofactor Modulating Catalase-Peroxidase Functions.
Li, J., Duan, R., Traore, E.S., Nguyen, R.C., Davis, I., Griffth, W.P., Goodwin, D.C., Jarzecki, A.A., Liu, A.(2024) Angew Chem Int Ed Engl 63: e202407018-e202407018
- PubMed: 39300819
- DOI: https://doi.org/10.1002/anie.202407018
- Primary Citation of Related Structures:
8CZP, 8U3P, 8W1W, 8W1X, 8W1Y - PubMed Abstract:
Bifunctional catalase-peroxidase (KatG) features a posttranslational methionine-tyrosine-tryptophan (MYW) crosslinked cofactor crucial for its catalase function, enabling pathogens to neutralize hydrogen peroxide during infection. We discovered the presence of indole nitrogen-linked hydroperoxyl adduct (MYW-OOH) in Mycobacterium tuberculosis KatG in the solution state under ambient conditions, suggesting its natural occurrence. By isolating predominantly MYW-OOH-containing KatG protein, we investigated the chemical stability and functional impact of MYW-OOH. We discovered that MYW-OOH inhibits catalase activity, presenting a unique temporary lock. Exposure to peroxide or increased temperature removes the hydroperoxyl adduct from the protein cofactor, converting MYW-OOH to MYW and restoring the detoxifying ability of the enzyme against hydrogen peroxide. Thus, the N-linked hydroperoxyl group is releasable. KatG with MYW-OOH represents a catalase dormant, but primed, state of the enzyme. These findings provide insight into chemical strategies targeting the bifunctional enzyme KatG in pathogens, highlighting the role of N-linked hydroperoxyl modifications in enzymatic function.
Organizational Affiliation:
University of Texas at San Antonio, Chemistry, UNITED STATES.