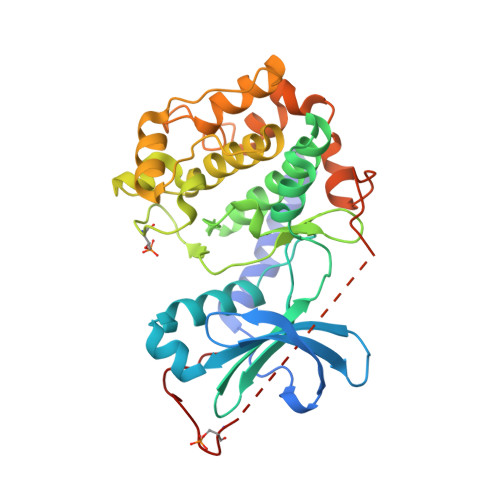
Structure and Function of the Human Sperm-Specific Isoform of Protein Kinase a (Pka) Catalytic Subunit Calpha2
Hereng, T.H., Backe, P.H., Kahmann, J., Scheich, C., Bjoras, M., Skalhegg, B.S., Rosendal, K.R.(2012) J Struct Biol 178: 300
- PubMed: 22504716
- DOI: https://doi.org/10.1016/j.jsb.2012.03.013
- Primary Citation of Related Structures:
4AE6, 4AE9 - PubMed Abstract:
Protein kinase A (PKA) exists as several tissue-specific isoforms that through phosphorylation of serine and threonine residues of substrate proteins act as key regulators of a number of cellular processes. We here demonstrate that the human sperm-specific isoform of PKA named Cα2 is important for sperm motility and thus male fertility. Furthermore, we report on the first three-dimensional crystal structure of human apo Cα2 to 2.1 Å. Apo Cα2 displays an open conformation similar to the well-characterized apo structure of murine Cα1. The asymmetric unit contains two molecules and the core of the small lobe is rotated by almost 13° in the A molecule relative to the B molecule. In addition, a salt bridge between Lys72 and Glu91 was observed for Cα2 in the apo-form, a conformation previously found only in dimeric or ternary complexes of Cα1. Human Cα2 and Cα1 share primary structure with the exception of the amino acids at the N-terminus coded for by an alternative exon 1. The N-terminal glycine of Cα1 is myristoylated and this aliphatic chain anchors the N-terminus to an intramolecular hydrophobic pocket. Cα2 cannot be myristoylated and the crystal structure revealed that the equivalent hydrophobic pocket is unoccupied and exposed. Nuclear magnetic resonance (NMR) spectroscopy further demonstrated that detergents with hydrophobic moieties of different lengths can bind deep into this uncovered pocket. Our findings indicate that Cα2 through the hydrophobic pocket has the ability to bind intracellular targets in the sperm cell, which may modulate protein stability, activity and/or cellular localization.
Organizational Affiliation:
Spermatech AS, N-0373 Oslo, Norway.