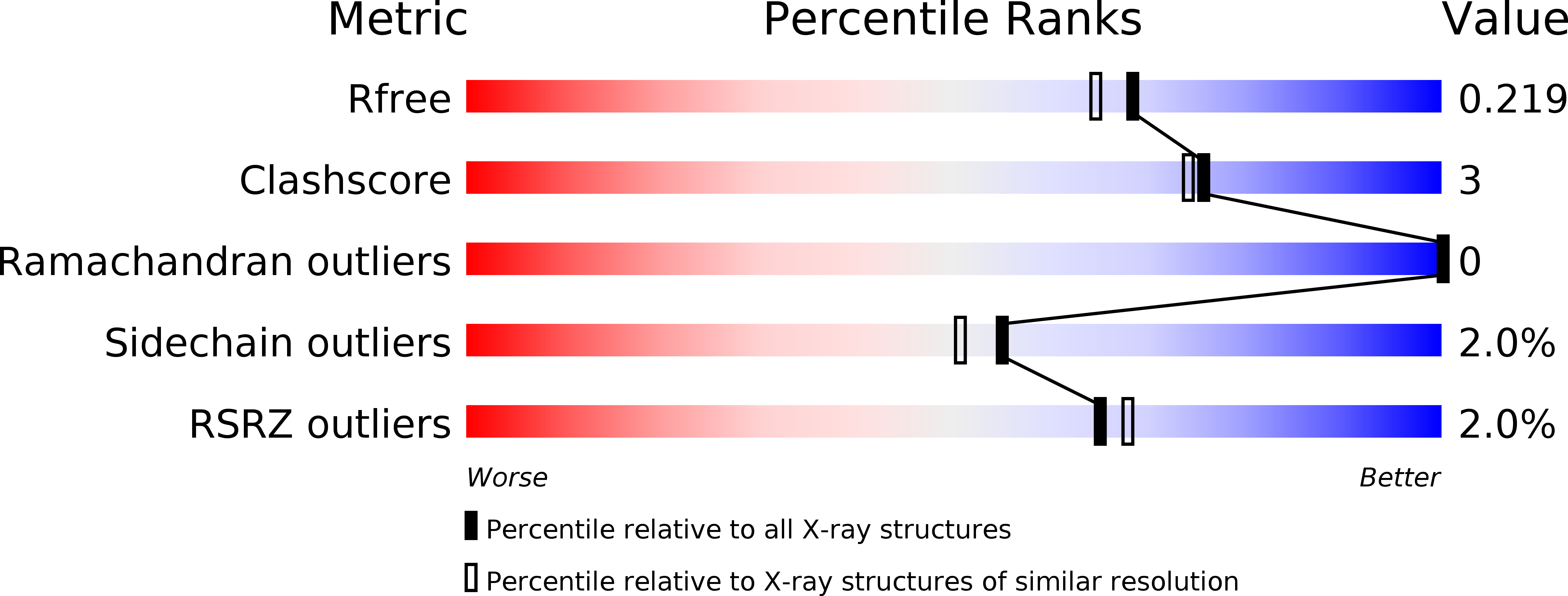
PARP Inhibitor with Selectivity Toward ADP-Ribosyltransferase ARTD3/PARP3
Lindgren, A.E.G., Karlberg, T., Thorsell, A.G., Hesse, M., Spjut, S., Ekblad, T., Andersson, C.D., Pinto, A.F., Weigelt, J., Hottiger, M.O., Linusson, A., Elofsson, M., Schuler, H.(2013) ACS Chem Biol 8: 1698-1703
- PubMed: 23742272
- DOI: https://doi.org/10.1021/cb4002014
- Primary Citation of Related Structures:
4GV0, 4GV2, 4GV4, 4GV7 - PubMed Abstract:
Inhibiting ADP-ribosyl transferases with PARP-inhibitors is considered a promising strategy for the treatment of many cancers and ischemia, but most of the cellular targets are poorly characterized. Here, we describe an inhibitor of ADP-ribosyltransferase-3/poly(ADP-ribose) polymerase-3 (ARTD3), a regulator of DNA repair and mitotic progression. In vitro profiling against 12 members of the enzyme family suggests selectivity for ARTD3, and crystal structures illustrate the molecular basis for inhibitor selectivity. The compound is active in cells, where it elicits ARTD3-specific effects at submicromolar concentration. Our results show that by targeting the nicotinamide binding site, selective inhibition can be achieved among the closest relatives of the validated clinical target, ADP-ribosyltransferase-1/poly(ADP-ribose) polymerase-1.
Organizational Affiliation:
Department of Chemistry, Umeå University, Umeå, Sweden.